[English] 日本語
 Yorodumi
Yorodumi- PDB-8efk: Structure of Lates calcarifer DNA polymerase theta polymerase dom... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8efk | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
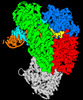
| Title | Structure of Lates calcarifer DNA polymerase theta polymerase domain with hairpin DNA | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / DNA double-strand break repair / Microhomology-mediated end joining / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology | 2',3'-dideoxyadenosine triphosphate / DNA / DNA (> 10) Function and homology information Function and homology information | |||||||||
| Biological species |  Lates calcarifer (barramundi perch) Lates calcarifer (barramundi perch) | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3 Å | |||||||||
 Authors Authors | Li, C. / Zhu, H. / Sun, J. / Gao, Y. | |||||||||
| Funding support |  United States, 2items United States, 2items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Structural basis of DNA polymerase θ mediated DNA end joining. Authors: Chuxuan Li / Hanwen Zhu / Shikai Jin / Leora M Maksoud / Nikhil Jain / Ji Sun / Yang Gao /  Abstract: DNA polymerase θ (Pol θ) plays an essential role in the microhomology-mediated end joining (MMEJ) pathway for repairing DNA double-strand breaks. However, the mechanisms by which Pol θ recognizes ...DNA polymerase θ (Pol θ) plays an essential role in the microhomology-mediated end joining (MMEJ) pathway for repairing DNA double-strand breaks. However, the mechanisms by which Pol θ recognizes microhomologous DNA ends and performs low-fidelity DNA synthesis remain unclear. Here, we present cryo-electron microscope structures of the polymerase domain of Lates calcarifer Pol θ with long and short duplex DNA at up to 2.4 Å resolution. Interestingly, Pol θ binds to long and short DNA substrates similarly, with extensive interactions around the active site. Moreover, Pol θ shares a similar active site as high-fidelity A-family polymerases with its finger domain well-closed but differs in having hydrophilic residues surrounding the nascent base pair. Computational simulations and mutagenesis studies suggest that the unique insertion loops of Pol θ help to stabilize short DNA binding and assemble the active site for MMEJ repair. Taken together, our results illustrate the structural basis of Pol θ-mediated MMEJ. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8efk.cif.gz 8efk.cif.gz | 135.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8efk.ent.gz pdb8efk.ent.gz | 96.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8efk.json.gz 8efk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  8efk_validation.pdf.gz 8efk_validation.pdf.gz | 1.2 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  8efk_full_validation.pdf.gz 8efk_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  8efk_validation.xml.gz 8efk_validation.xml.gz | 29.4 KB | Display | |
| Data in CIF |  8efk_validation.cif.gz 8efk_validation.cif.gz | 41 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ef/8efk https://data.pdbj.org/pub/pdb/validation_reports/ef/8efk ftp://data.pdbj.org/pub/pdb/validation_reports/ef/8efk ftp://data.pdbj.org/pub/pdb/validation_reports/ef/8efk | HTTPS FTP |
-Related structure data
| Related structure data | 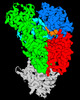 28084MC  8ef9C  8efcC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 95674.117 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Lates calcarifer (barramundi perch) Lates calcarifer (barramundi perch)Production host:  |
|---|---|
| #2: DNA chain | Mass: 7278.689 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Lates calcarifer (barramundi perch) Lates calcarifer (barramundi perch) |
| #3: Chemical | ChemComp-MG / |
| #4: Chemical | ChemComp-DDS / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Ternary complex of Lates calcarifer DNA polymerase Theta with duplex DNA and incoming nucleotide Type: COMPLEX / Entity ID: #1-#2 / Source: MULTIPLE SOURCES |
|---|---|
| Source (natural) | Organism:  Lates calcarifer (barramundi perch) Lates calcarifer (barramundi perch) |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 7.6 |
| Specimen | Conc.: 0.8 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 295 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 1600 nm / Nominal defocus min: 800 nm |
| Image recording | Electron dose: 1.2 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.19.2_4158: / Classification: refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 288920 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj