[English] 日本語
 Yorodumi
Yorodumi- PDB-5k2f: Structure of NNQQNY from yeast prion Sup35 with cadmium acetate d... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5k2f | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of NNQQNY from yeast prion Sup35 with cadmium acetate determined by MicroED | ||||||||||||||||||
 Components Components | Eukaryotic peptide chain release factor GTP-binding subunit | ||||||||||||||||||
 Keywords Keywords | PROTEIN FIBRIL / amyloid / yeast prion | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationEukaryotic Translation Termination / translation release factor complex / translation release factor activity / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / translational termination / cytoplasmic stress granule / regulation of translation / ribosome binding ...Eukaryotic Translation Termination / translation release factor complex / translation release factor activity / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / translational termination / cytoplasmic stress granule / regulation of translation / ribosome binding / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / translation / mRNA binding / GTPase activity / GTP binding / identical protein binding / cytosol / cytoplasm Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | ELECTRON CRYSTALLOGRAPHY / electron crystallography / cryo EM / Resolution: 1 Å | ||||||||||||||||||
 Authors Authors | Rodriguez, J.A. / Sawaya, M.R. / Cascio, D. / Eisenberg, D.S. | ||||||||||||||||||
| Funding support |  United States, 5items United States, 5items
| ||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2016 Journal: Proc Natl Acad Sci U S A / Year: 2016Title: Ab initio structure determination from prion nanocrystals at atomic resolution by MicroED. Authors: Michael R Sawaya / Jose Rodriguez / Duilio Cascio / Michael J Collazo / Dan Shi / Francis E Reyes / Johan Hattne / Tamir Gonen / David S Eisenberg /  Abstract: Electrons, because of their strong interaction with matter, produce high-resolution diffraction patterns from tiny 3D crystals only a few hundred nanometers thick in a frozen-hydrated state. This ...Electrons, because of their strong interaction with matter, produce high-resolution diffraction patterns from tiny 3D crystals only a few hundred nanometers thick in a frozen-hydrated state. This discovery offers the prospect of facile structure determination of complex biological macromolecules, which cannot be coaxed to form crystals large enough for conventional crystallography or cannot easily be produced in sufficient quantities. Two potential obstacles stand in the way. The first is a phenomenon known as dynamical scattering, in which multiple scattering events scramble the recorded electron diffraction intensities so that they are no longer informative of the crystallized molecule. The second obstacle is the lack of a proven means of de novo phase determination, as is required if the molecule crystallized is insufficiently similar to one that has been previously determined. We show with four structures of the amyloid core of the Sup35 prion protein that, if the diffraction resolution is high enough, sufficiently accurate phases can be obtained by direct methods with the cryo-EM method microelectron diffraction (MicroED), just as in X-ray diffraction. The success of these four experiments dispels the concern that dynamical scattering is an obstacle to ab initio phasing by MicroED and suggests that structures of novel macromolecules can also be determined by direct methods. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5k2f.cif.gz 5k2f.cif.gz | 18.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5k2f.ent.gz pdb5k2f.ent.gz | 8.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5k2f.json.gz 5k2f.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/k2/5k2f https://data.pdbj.org/pub/pdb/validation_reports/k2/5k2f ftp://data.pdbj.org/pub/pdb/validation_reports/k2/5k2f ftp://data.pdbj.org/pub/pdb/validation_reports/k2/5k2f | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  8197MC  8196C  8198C 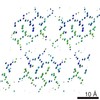 8199C  5k2eC  5k2gC 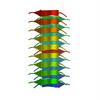 5k2hC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological unit is an extended pair of beta sheets comprising peptides at positions X,Y,Z and -X,Y+1/2,-Z extended ad infinitum along the b crystal axis. |
- Components
Components
| #1: Protein/peptide | Mass: 779.755 Da / Num. of mol.: 1 / Fragment: UNP residues 8-13 / Source method: obtained synthetically / Source: (synth.)  |
|---|---|
| #2: Chemical | ChemComp-CD / |
| #3: Chemical | ChemComp-ACT / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON CRYSTALLOGRAPHY |
|---|---|
| EM experiment | Aggregation state: 3D ARRAY / 3D reconstruction method: electron crystallography |
- Sample preparation
Sample preparation
| Component | Name: Prion fibril composed of a 6-residue segment of Sup35 and cadmium Type: COMPLEX / Entity ID: #1 / Source: MULTIPLE SOURCES | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular weight | Value: 3.25 kDa/nm / Experimental value: NO | ||||||||||||||||||||
| Source (natural) | Organism:  | ||||||||||||||||||||
| EM crystal formation | Instrument: 24-well plate Atmosphere: in air, in sealed chamber, in equilibrium with reservoir solution Details: Crystals were grown by hanging-drop vapor diffusion at ~20 degrees C. The reservoir solution contained 100 mM HEPES, pH 7.0, and 1 M sodium acetate (pH not adjusted). Drops were prepared by ...Details: Crystals were grown by hanging-drop vapor diffusion at ~20 degrees C. The reservoir solution contained 100 mM HEPES, pH 7.0, and 1 M sodium acetate (pH not adjusted). Drops were prepared by pipetting 5 microliters of 30 mg/mL aqueous peptide solution, 4 microliters of reservoir solution, and 1 microliter of 0.1 M zinc sulfate onto a glass coverslip. Crystals grew within a day. Lipid mixture: none / Temperature: 298 K / Time: 1 DAY | ||||||||||||||||||||
| Buffer solution | pH: 7 | ||||||||||||||||||||
| Buffer component |
| ||||||||||||||||||||
| Specimen | Conc.: 30 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES / Details: crystal | ||||||||||||||||||||
| Specimen support | Grid material: COPPER / Grid mesh size: 300 divisions/in. / Grid type: Quantifoil R2/2 | ||||||||||||||||||||
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Details: Plunged into liquid ethane (FEI VITROBOT MARK IV) | ||||||||||||||||||||
| Crystal grow | Temperature: 273 K / Method: vapor diffusion, hanging drop / pH: 7 Details: 1.0 M sodium acetate, 0.1 M HEPES, pH 7.0, 0.01 M cadmium sulfate |
-Data collection
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Microscopy | Model: FEI TECNAI F20 | ||||||||||||||||||||||||||||||||||||||||||
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM | ||||||||||||||||||||||||||||||||||||||||||
| Electron lens | Mode: DIFFRACTION / Alignment procedure: BASIC | ||||||||||||||||||||||||||||||||||||||||||
| Specimen holder | Cryogen: NITROGEN Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Temperature (max): 100 K / Temperature (min): 100 K | ||||||||||||||||||||||||||||||||||||||||||
| Image recording | Average exposure time: 2 sec. / Electron dose: 0.01 e/Å2 / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Num. of diffraction images: 650 / Num. of grids imaged: 2 Details: The detector was operated in rolling shutter mode with 2x2 pixel binning. | ||||||||||||||||||||||||||||||||||||||||||
| Image scans | Sampling size: 15.6 µm / Width: 4096 / Height: 4096 | ||||||||||||||||||||||||||||||||||||||||||
| EM diffraction | Camera length: 1350 mm | ||||||||||||||||||||||||||||||||||||||||||
| EM diffraction shell |
| ||||||||||||||||||||||||||||||||||||||||||
| EM diffraction stats | Details: Phase statistics are not applicable. No imaging was used. The phases were obtained by a crystallographic direct methods program, SHELXD. Fourier space coverage: 82.7 % / High resolution: 1 Å / Num. of intensities measured: 16753 / Num. of structure factors: 2399 / Phase error: 0.1 ° / Phase residual: 0.1 ° / Phase error rejection criteria: 0 / Rmerge: 15.1 / Rsym: 15.1 | ||||||||||||||||||||||||||||||||||||||||||
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||
| Diffraction source | Source: ELECTRON MICROSCOPE / Type: TECNAI F20 TEM / Wavelength: 0.0251 Å | ||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: TVIPS F416 CMOS CAMERA / Detector: CMOS / Date: Feb 3, 2016 | ||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.0251 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1→90 Å / Num. obs: 2564 / % possible obs: 84.5 % / Redundancy: 4.6 % / Rmerge(I) obs: 0.18 / Χ2: 1.222 / Net I/av σ(I): 7.346 / Net I/σ(I): 5.1 / Num. measured all: 11810 | ||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1 / Rejects: _
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EM 3D crystal entity | ∠α: 90 ° / ∠β: 104.29 ° / ∠γ: 90 ° / A: 22.074 Å / B: 4.932 Å / C: 23.51 Å / Space group name: P1211 / Space group num: 4 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTF correction | Type: NONE | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution method: DIFFRACTION PATTERN/LAYERLINES Details: The density map was obtained using measured diffraction intensities and phases acquired from crystallographic direct methods program SHELXD. Symmetry type: 3D CRYSTAL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | B value: 6.44 / Protocol: OTHER / Space: RECIPROCAL / Target criteria: maximum likelihood | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | Resolution: 1→22.79 Å / Cor.coef. Fo:Fc: 0.922 / Cor.coef. Fo:Fc free: 0.934 / SU B: 2.367 / SU ML: 0.048 / SU R Cruickshank DPI: 0.0475 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.047 / ESU R Free: 0.045 / Details: HYDROGENS HAVE BEEN USED IF PRESENT IN THE INPUT
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 22.93 Å2 / Biso mean: 6.444 Å2 / Biso min: 3.46 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 0.98→22.79 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 0.978→1.004 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller








 PDBj
PDBj












