+ Open data
Open data
- Basic information
Basic information
| Entry | 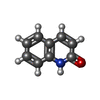 Database: PDB chemical components / ID: OCH Database: PDB chemical components / ID: OCH |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: OCH / Model coordinates PDB-ID: 1Z03 | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda Brenda     / /  ChEBI ChEBI  / /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  PubChem / PubChem /  SureChEMBL SureChEMBL    / /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 6 items

PDB-1z03: 
2-Oxoquinoline 8-Monooxygenase Component: Active site Modulation by Rieske-[2fe-2S] Center Oxidation/Reduction

PDB-3srg: 
Serum paraoxonase-1 by directed evolution at pH 6.5 in complex with 2-hydroxyquinoline

PDB-7hiz: 
PanDDA analysis group deposition -- Crystal structure of ABLE in complex with ZINC000008579298

PDB-7riz: 
Crystal structure of RPA3624, a beta-propeller lactonase from Rhodopseudomonas palustris, with active-site bound 2-hydroxyquinoline

PDB-9idm: 
Human Deoxyhypusine Synthase Fragment Screening Campaign - ligand VT00175

PDB-9r0q: 
Paraoxonase-1 in complex with terbium(III) and 2-hydroxyquinoline
 Movie
Movie Controller
Controller



