 Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | 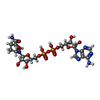 Database: PDB chemical components / ID: NAX Database: PDB chemical components / ID: NAX |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: NAX / Model coordinates PDB-ID: 1BDM | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda Brenda   / /  ChEBI / ChEBI /  HMDB / HMDB /  PubChem / PubChem /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| OpenEye OEToolkits 1.5.0 | [[( |
|---|
-PDB entries
Showing all 5 items

PDB-1bdm: 
THE STRUCTURE AT 1.8 ANGSTROMS RESOLUTION OF A SINGLE SITE MUTANT (T189I) OF MALATE DEHYDROGENASE FROM THERMUS FLAVUS WITH INCREASED ENZYMATIC ACTIVITY

PDB-3rq2: 
Crystal Structure of ADP/ATP-dependent NAD(P)H-hydrate dehydratase from Bacillus subtilis co-crystallized with ATP/Mg2+ and soaked with NADH

PDB-3rsq: 
Crystal structure of tm0922, a fusion of a domain of unknown function and ADP/ATP-dependent NAD(P)H-hydrate dehydratase from Thermotoga maritima soaked with NADH

PDB-3rtd: 
Crystal structure of tm0922, a fusion of a domain of unknown function and ADP/ATP-dependent NAD(P)H-hydrate dehydratase from Thermotoga maritima soaked with NADH and ADP.

PDB-6dbb: 
Crystal structure of a Putative aldehyde dehydrogenase family protein Burkholderia cenocepacia J2315 in complex with partially reduced NADH
 Movie
Movie Controller
Controller


