+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9541 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of TRiC-AMP-PNP | |||||||||||||||
 Map data Map data | Yeast TRiC-AMP-PNP at 4.6 angstroms resolution | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Chaperonin / Yeast / CHAPERONE | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationAssociation of TriC/CCT with target proteins during biosynthesis / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / chaperonin-containing T-complex / Neutrophil degranulation / ATP-dependent protein folding chaperone / : / protein folding / ATP hydrolysis activity / ATP binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.6 Å | |||||||||||||||
 Authors Authors | Zang Y / Jin M | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2016 Journal: Nat Struct Mol Biol / Year: 2016Title: Staggered ATP binding mechanism of eukaryotic chaperonin TRiC (CCT) revealed through high-resolution cryo-EM. Authors: Yunxiang Zang / Mingliang Jin / Huping Wang / Zhicheng Cui / Liangliang Kong / Caixuan Liu / Yao Cong /  Abstract: The eukaryotic chaperonin TRiC (or CCT) assists in the folding of 10% of cytosolic proteins. Here we present two cryo-EM structures of Saccharomyces cerevisiae TRiC in a newly identified nucleotide ...The eukaryotic chaperonin TRiC (or CCT) assists in the folding of 10% of cytosolic proteins. Here we present two cryo-EM structures of Saccharomyces cerevisiae TRiC in a newly identified nucleotide partially preloaded (NPP) state and in the ATP-bound state, at 4.7-Å and 4.6-Å resolution, respectively. Through inner-subunit eGFP tagging, we identified the subunit locations in open-state TRiC and found that the CCT2 subunit pair forms an unexpected Z shape. ATP binding induces a dramatic conformational change on the CCT2 side, thereby suggesting that CCT2 plays an essential role in TRiC allosteric cooperativity. Our structural and biochemical data reveal a staggered ATP binding mechanism of TRiC with preloaded nucleotide on the CCT6 side of NPP-TRiC and demonstrate that TRiC has evolved into a complex that is structurally divided into two sides. This work offers insight into how the TRiC nucleotide cycle coordinates with its mechanical cycle in preparing folding intermediates for further productive folding. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9541.map.gz emd_9541.map.gz | 7.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9541-v30.xml emd-9541-v30.xml emd-9541.xml emd-9541.xml | 24.1 KB 24.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9541.png emd_9541.png | 184 KB | ||
| Filedesc metadata |  emd-9541.cif.gz emd-9541.cif.gz | 8.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9541 http://ftp.pdbj.org/pub/emdb/structures/EMD-9541 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9541 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9541 | HTTPS FTP |
-Related structure data
| Related structure data |  5gw5MC  9540C  5gw4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9541.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9541.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Yeast TRiC-AMP-PNP at 4.6 angstroms resolution | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.318 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
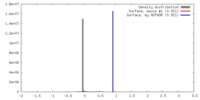
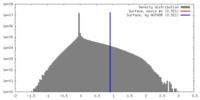
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Yeast TRiC-AMP-PNP
+Supramolecule #1: Yeast TRiC-AMP-PNP
+Supramolecule #2: T-complex protein 1 subunit alpha
+Supramolecule #3: T-complex protein 1 subunit beta
+Supramolecule #4: T-complex protein 1 subunit delta
+Supramolecule #5: T-complex protein 1 subunit epsilon
+Supramolecule #6: T-complex protein 1 subunit gamma
+Supramolecule #7: T-complex protein 1 subunit eta
+Supramolecule #8: T-complex protein 1 subunit theta
+Supramolecule #9: T-complex protein 1 subunit zeta
+Macromolecule #1: T-complex protein 1 subunit alpha
+Macromolecule #2: T-complex protein 1 subunit beta
+Macromolecule #3: T-complex protein 1 subunit delta
+Macromolecule #4: T-complex protein 1 subunit epsilon
+Macromolecule #5: T-complex protein 1 subunit gamma
+Macromolecule #6: T-complex protein 1 subunit eta
+Macromolecule #7: T-complex protein 1 subunit theta
+Macromolecule #8: T-complex protein 1 subunit zeta
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 10 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 3 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 7.6 sec. / Average electron dose: 38.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.0 µm / Calibrated defocus min: 1.5 µm / Calibrated magnification: 18000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.005 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-5gw5: |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)