[English] 日本語
 Yorodumi
Yorodumi- EMDB-9538: Kinesin-8 motor, KIF19A, in the nucleotide-free state complexed w... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9538 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
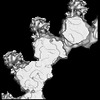
| Title | Kinesin-8 motor, KIF19A, in the nucleotide-free state complexed with GDP-taxol microtubule | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinesin-8 / KIF19A / GDP-taxol-microtubule / Plus-end directed motor / Microtubule depolymerization / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationaxonemal microtubule depolymerization / plus-end specific microtubule depolymerization / Kinesins / COPI-dependent Golgi-to-ER retrograde traffic / MHC class II antigen presentation / plus-end-directed microtubule motor activity / microtubule motor activity / kinesin complex / microtubule-based movement / axoneme ...axonemal microtubule depolymerization / plus-end specific microtubule depolymerization / Kinesins / COPI-dependent Golgi-to-ER retrograde traffic / MHC class II antigen presentation / plus-end-directed microtubule motor activity / microtubule motor activity / kinesin complex / microtubule-based movement / axoneme / microtubule binding / microtubule / cilium / ATP hydrolysis activity / ATP binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 7.0 Å | |||||||||
 Authors Authors | Morikawa M / Nitta R / Yajima H / Shigematsu H / Kikkawa M / Hirokawa N | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2016 Journal: Elife / Year: 2016Title: Motility and microtubule depolymerization mechanisms of the Kinesin-8 motor, KIF19A. Authors: Doudou Wang / Ryo Nitta / Manatsu Morikawa / Hiroaki Yajima / Shigeyuki Inoue / Hideki Shigematsu / Masahide Kikkawa / Nobutaka Hirokawa /   Abstract: The kinesin-8 motor, KIF19A, accumulates at cilia tips and controls cilium length. Defective KIF19A leads to hydrocephalus and female infertility because of abnormally elongated cilia. Uniquely among ...The kinesin-8 motor, KIF19A, accumulates at cilia tips and controls cilium length. Defective KIF19A leads to hydrocephalus and female infertility because of abnormally elongated cilia. Uniquely among kinesins, KIF19A possesses the dual functions of motility along ciliary microtubules and depolymerization of microtubules. To elucidate the molecular mechanisms of these functions we solved the crystal structure of its motor domain and determined its cryo-electron microscopy structure complexed with a microtubule. The features of KIF19A that enable its dual function are clustered on its microtubule-binding side. Unexpectedly, a destabilized switch II coordinates with a destabilized L8 to enable KIF19A to adjust to both straight and curved microtubule protofilaments. The basic clusters of L2 and L12 tether the microtubule. The long L2 with a characteristic acidic-hydrophobic-basic sequence effectively stabilizes the curved conformation of microtubule ends. Hence, KIF19A utilizes multiple strategies to accomplish the dual functions of motility and microtubule depolymerization by ATP hydrolysis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9538.map.gz emd_9538.map.gz | 10.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9538-v30.xml emd-9538-v30.xml emd-9538.xml emd-9538.xml | 10 KB 10 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9538.png emd_9538.png | 188.5 KB | ||
| Filedesc metadata |  emd-9538.cif.gz emd-9538.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9538 http://ftp.pdbj.org/pub/emdb/structures/EMD-9538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9538 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9538 | HTTPS FTP |
-Related structure data
| Related structure data |  5gsyMC  5gszC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9538.map.gz / Format: CCP4 / Size: 12 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9538.map.gz / Format: CCP4 / Size: 12 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.284 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
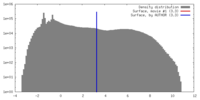
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : KIF19A motor domain complexed with GDP-taxol-MT
| Entire | Name: KIF19A motor domain complexed with GDP-taxol-MT |
|---|---|
| Components |
|
-Supramolecule #1: KIF19A motor domain complexed with GDP-taxol-MT
| Supramolecule | Name: KIF19A motor domain complexed with GDP-taxol-MT / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Kinesin-like protein KIF19
| Macromolecule | Name: Kinesin-like protein KIF19 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.636031 KDa |
| Recombinant expression | Organism: |
| Sequence | String: MKDSGDSKDQ QLMVALRVRP ISVAELEEGA TLIAHKMDEQ MVVLMDPMED PDDILRAHRS REKSYLFDVA FDFTATQEMV YQATTKSLI EGVISGYNAT VFAYGPTGCG KTYTMLGTDH EPGIYVRTLN DLFRAIEETS NDMEYEVSMS YLEIYNEMIR D LLNPALGY ...String: MKDSGDSKDQ QLMVALRVRP ISVAELEEGA TLIAHKMDEQ MVVLMDPMED PDDILRAHRS REKSYLFDVA FDFTATQEMV YQATTKSLI EGVISGYNAT VFAYGPTGCG KTYTMLGTDH EPGIYVRTLN DLFRAIEETS NDMEYEVSMS YLEIYNEMIR D LLNPALGY LELREDSKGV IQVAGITEVS TINAKEIMQL LMKGNRQRTQ EPTAANQTSS RSHAVLQVAV RQRSRVKNIL QE VRQGRLF MIDLAGSERA SQTQNRGQRM KEGAHINRSL LALGNCINAL SDKGSNKYIN YRDSKLTRLL KDSLGGNSRT VMI AHISPA STAFEESRNT LTYAGRAKNI RTRVKQNHHH HHHH UniProtKB: Kinesin-like protein KIF19 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 200 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 300 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 8.727751 Å Applied symmetry - Helical parameters - Δ&Phi: -25.717620 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 7.0 Å / Resolution method: FSC 0.143 CUT-OFF / Details: High-resolution noise substitution was performed. / Number images used: 471380 |
|---|---|
| Startup model | Type of model: PDB ENTRY |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)