+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9378 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | PGT145 Fab in complex with full length AMC011 HIV-1 Env | ||||||||||||||||||
 Map data Map data | PGT145 Fab in complex with full length AMC011 HIV-1 Env | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.7 Å | ||||||||||||||||||
 Authors Authors | Torrents de la Pena A / Rantalainen K / Torres JL / Ward AB | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2019 Journal: PLoS Pathog / Year: 2019Title: Similarities and differences between native HIV-1 envelope glycoprotein trimers and stabilized soluble trimer mimetics. Authors: Alba Torrents de la Peña / Kimmo Rantalainen / Christopher A Cottrell / Joel D Allen / Marit J van Gils / Jonathan L Torres / Max Crispin / Rogier W Sanders / Andrew B Ward /    Abstract: The HIV-1 envelope glycoprotein (Env) trimer is located on the surface of the virus and is the target of broadly neutralizing antibodies (bNAbs). Recombinant native-like soluble Env trimer mimetics, ...The HIV-1 envelope glycoprotein (Env) trimer is located on the surface of the virus and is the target of broadly neutralizing antibodies (bNAbs). Recombinant native-like soluble Env trimer mimetics, such as SOSIP trimers, have taken a central role in HIV-1 vaccine research aimed at inducing bNAbs. We therefore performed a direct and thorough comparison of a full-length unmodified Env trimer containing the transmembrane domain and the cytoplasmic tail, with the sequence matched soluble SOSIP trimer, both based on an early Env sequence (AMC011) from an HIV+ individual that developed bNAbs. The structures of the full-length AMC011 trimer bound to either bNAb PGT145 or PGT151 were very similar to the structures of SOSIP trimers. Antigenically, the full-length and SOSIP trimers were comparable, but in contrast to the full-length trimer, the SOSIP trimer did not bind at all to non-neutralizing antibodies, most likely as a consequence of the intrinsic stabilization of the SOSIP trimer. Furthermore, the glycan composition of full-length and SOSIP trimers was similar overall, but the SOSIP trimer possessed slightly less complex and less extensively processed glycans, which may relate to the intrinsic stabilization as well as the absence of the membrane tether. These data provide insights into how to best use and improve membrane-associated full-length and soluble SOSIP HIV-1 Env trimers as immunogens. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9378.map.gz emd_9378.map.gz | 117.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9378-v30.xml emd-9378-v30.xml emd-9378.xml emd-9378.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9378.png emd_9378.png | 31.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9378 http://ftp.pdbj.org/pub/emdb/structures/EMD-9378 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9378 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9378 | HTTPS FTP |
-Validation report
| Summary document |  emd_9378_validation.pdf.gz emd_9378_validation.pdf.gz | 492.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9378_full_validation.pdf.gz emd_9378_full_validation.pdf.gz | 492.4 KB | Display | |
| Data in XML |  emd_9378_validation.xml.gz emd_9378_validation.xml.gz | 6.4 KB | Display | |
| Data in CIF |  emd_9378_validation.cif.gz emd_9378_validation.cif.gz | 7.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9378 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9378 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9378 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9378 | HTTPS FTP |
-Related structure data
| Related structure data |  6nijMC  6olpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9378.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9378.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PGT145 Fab in complex with full length AMC011 HIV-1 Env | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
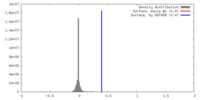
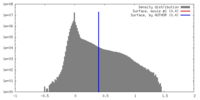
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PGT145 Fab in complex with full length AMC011 HIV-1 Env
| Entire | Name: PGT145 Fab in complex with full length AMC011 HIV-1 Env |
|---|---|
| Components |
|
-Supramolecule #1: PGT145 Fab in complex with full length AMC011 HIV-1 Env
| Supramolecule | Name: PGT145 Fab in complex with full length AMC011 HIV-1 Env type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: PGT145
| Supramolecule | Name: PGT145 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: AMC011 Glycoprotein 120
| Supramolecule | Name: AMC011 Glycoprotein 120 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: AMC011 Glycoprotein 41
| Supramolecule | Name: AMC011 Glycoprotein 41 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: PGT145 Fab heavy chain
| Macromolecule | Name: PGT145 Fab heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.550196 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGAE VKKPGSSVKV SCKASGNSFS NHDVHWVRQA TGQGLEWMGW MSHEGDKTGL AQKFQGRVTI TRDSGASTVY MELRGLTAD DTAIYYCLTG SKHRLRDYFL YNEYGPNYEE WGDYLATLDV WGHGTAVTVS S |
-Macromolecule #2: PGT145 Fab light chain
| Macromolecule | Name: PGT145 Fab light chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.350922 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVVITQSPLF LPVTPGEAAS LSCKCSHSLQ HSTGANYLAW YLQRPGQTPR LLIHLATHRA SGVPDRFSGS GSGTDFTLKI SRVESDDVG TYYCMQGLHS PWTFGQGTKV EIKR |
-Macromolecule #3: AMC011 Glycoprotein 120
| Macromolecule | Name: AMC011 Glycoprotein 120 / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 52.880039 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AEQLWVTVYY GVPVWKEATT TLFCASDARA YDTEVHNVWA THACVPTDPN PQEVVLENVT ENFNMWKNNM VEQMHEDIIS LWDQSLKPC VKLTPLCVTL NCTDLRNATN TNATNTTSSS RGTMEGGEIK NCSFNITTSM RDKVQKEYAL FYKLDVVPIK N DNTSYRLI ...String: AEQLWVTVYY GVPVWKEATT TLFCASDARA YDTEVHNVWA THACVPTDPN PQEVVLENVT ENFNMWKNNM VEQMHEDIIS LWDQSLKPC VKLTPLCVTL NCTDLRNATN TNATNTTSSS RGTMEGGEIK NCSFNITTSM RDKVQKEYAL FYKLDVVPIK N DNTSYRLI SCNTSVITQA CPKVSFEPIP IHYCAPAGFA ILKCNDKKFN GTGPCTNVST VQCTHGIRPV VSTQLLLNGS LA EEEVVIR SANFTDNAKI IIVQLNKSVE INCTRPNNNT RKSIHIGPGR AFYTTGEIIG DIRQAHCNIS GTKWNDTLKQ IVV KLKEQF GNKTIVFNHS SGGDPEIVMH SFNCGGEFFY CNSTQLFNST WNDTTGSNYT GTIVLPCRIK QIVNMWQEVG KAMY APPIK GQIRCSSNIT GLILIRDGGK NRSENTEIFR PGGGDMRDNW RSELYKYKVV KIEPLGIAPT KAKRRVVQ |
-Macromolecule #4: AMC011 Glycoprotein 41
| Macromolecule | Name: AMC011 Glycoprotein 41 / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 39.677762 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RLLLSGIVQQ QNNLLRAIEA QQHLLQLTVW GIKQLQARVL AVERYLKDQQ LLGIWGCSG KLICTTAVPW NTSWSNKSYN QIWNNMTWME WEREIDNYTS LIYTLIEDSQ NQQEKNEQEL LELDKWASLW N WFDITKWL ...String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RLLLSGIVQQ QNNLLRAIEA QQHLLQLTVW GIKQLQARVL AVERYLKDQQ LLGIWGCSG KLICTTAVPW NTSWSNKSYN QIWNNMTWME WEREIDNYTS LIYTLIEDSQ NQQEKNEQEL LELDKWASLW N WFDITKWL WYIKIFIMIV GGLIGLRIVF TVLSIVNRIR QGYSPLSFQT PLPTPRGPDR PEGIEEEGGE RDRDRSDRLV TG FLALIWV DLRSLCLFSY HRLRDLLLIV TRIVELLGRR GWGVLKYWWN LLQYWSQELR NSAVSLLNAT AIAVAEGTDR VIE VSQRAF RAILHVPVRI RQGLERALV |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 5 sec. |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: OTHER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 5.7 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 51588 |
|---|---|
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6nij: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)