+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of GGCX-MGP complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Vitamin K-dependent gamma-carboxylase binding to its substrate MGP / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationstructural constituent of bone / peptidyl-glutamate 4-carboxylase / gamma-glutamyl carboxylase activity / negative regulation of testosterone biosynthetic process / negative regulation of bone development / Defective gamma-carboxylation of F9 / vitamin binding / vitamin K metabolic process / negative regulation of neurotransmitter secretion / regulation of bone mineralization ...structural constituent of bone / peptidyl-glutamate 4-carboxylase / gamma-glutamyl carboxylase activity / negative regulation of testosterone biosynthetic process / negative regulation of bone development / Defective gamma-carboxylation of F9 / vitamin binding / vitamin K metabolic process / negative regulation of neurotransmitter secretion / regulation of bone mineralization / type B pancreatic cell proliferation / cartilage condensation / extracellular matrix structural constituent / Gamma-carboxylation of protein precursors / ossification / protein modification process / protein maturation / cellular response to insulin stimulus / blood coagulation / glucose homeostasis / extracellular matrix / cell differentiation / endoplasmic reticulum lumen / calcium ion binding / endoplasmic reticulum membrane / extracellular exosome / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.33 Å | |||||||||
 Authors Authors | Qian HW / Zhang WJ | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of GGcX-MGP complex Authors: Qian HW / Zhang WJ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_65925.map.gz emd_65925.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-65925-v30.xml emd-65925-v30.xml emd-65925.xml emd-65925.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_65925.png emd_65925.png | 54.6 KB | ||
| Filedesc metadata |  emd-65925.cif.gz emd-65925.cif.gz | 6.2 KB | ||
| Others |  emd_65925_half_map_1.map.gz emd_65925_half_map_1.map.gz emd_65925_half_map_2.map.gz emd_65925_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-65925 http://ftp.pdbj.org/pub/emdb/structures/EMD-65925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-65925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-65925 | HTTPS FTP |
-Related structure data
| Related structure data |  9wfcMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_65925.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_65925.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_65925_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_65925_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of complex D
| Entire | Name: Cryo-EM structure of complex D |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of complex D
| Supramolecule | Name: Cryo-EM structure of complex D / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Vitamin K-dependent gamma-carboxylase
| Macromolecule | Name: Vitamin K-dependent gamma-carboxylase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidyl-glutamate 4-carboxylase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 81.275875 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SRIGKLLGFE WTDLSSWRRL VTLLNRPTDP ASLAVFRFLF GFLMVLDIPQ ERGLSSLDRK YLDGLDVCRF PLLDALRPLP LDWMYLVYT IMFLGALGMM LGLCYRISCV LFLLPYWYVF LLDKTSWNNH SYLYGLLAFQ LTFMDANHYW SVDGLLNAHR R NAHVPLWN ...String: SRIGKLLGFE WTDLSSWRRL VTLLNRPTDP ASLAVFRFLF GFLMVLDIPQ ERGLSSLDRK YLDGLDVCRF PLLDALRPLP LDWMYLVYT IMFLGALGMM LGLCYRISCV LFLLPYWYVF LLDKTSWNNH SYLYGLLAFQ LTFMDANHYW SVDGLLNAHR R NAHVPLWN YAVLRGQIFI VYFIAGVKKL DADWVEGYSM EYLSRHWLFS PFKLLLSEEL TSLLVVHWGG LLLDLSAGFL LF FDVSRSI GLFFVSYFHC MNSQLFSIGM FSYVMLASSP LFCSPEWPRK LVSYCPRRLQ QLLPLKAAPQ PSVSCVYKRS RGK SGQKPG LRHQLGAAFT LLYLLEQLFL PYSHFLTQGY NNWTNGLYGY SWDMMVHSRS HQHVKITYRD GRTGELGYLN PGVF TQSRR WKDHADMLKQ YATCLSRLLP KYNVTEPQIY FDIWVSINDR FQQRIFDPRV DIVQAAWSPF QRTSWVQPLL MDLSP WRAK LQEIKSSLDN HTEVVFIADF PGLHLENFVS EDLGNTSIQL LQGEVTVELV AEQKNQTLRE GEKMQLPAGE YHKVYT TSP SPSCYMYVYV NTTELALEQD LAYLQELKEK VENGSETGPL PPELQPLLEG EVKGGPEPTP LVQTFLRRQQ RLQEIER RR NTPFHERFFR FLLRKLYVFR RSFLMTCISL RNLILGRPSL EQLAQEVTYA NLRPF UniProtKB: Vitamin K-dependent gamma-carboxylase |
-Macromolecule #2: Matrix Gla protein
| Macromolecule | Name: Matrix Gla protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.336689 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LCYESHESMS YELNPFINRR NANTFISP UniProtKB: Matrix Gla protein |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 5 / Number of copies: 2 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #6: (2R)-3-(phosphonooxy)propane-1,2-diyl (9Z,9'Z)bis-octadec-9-enoate
| Macromolecule | Name: (2R)-3-(phosphonooxy)propane-1,2-diyl (9Z,9'Z)bis-octadec-9-enoate type: ligand / ID: 6 / Number of copies: 1 / Formula: MX7 |
|---|---|
| Molecular weight | Theoretical: 700.966 Da |
| Chemical component information | 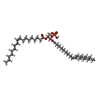 ChemComp-MX7: |
-Macromolecule #7: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 7 / Number of copies: 1 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #8: (1aR,7aS)-1a-methyl-7a-[(2E,6E,10E)-3,7,11,15-tetramethylhexadeca...
| Macromolecule | Name: (1aR,7aS)-1a-methyl-7a-[(2E,6E,10E)-3,7,11,15-tetramethylhexadeca-2,6,10,14-tetraen-1-yl]-1a,7a-dihydronaphtho[2,3-b]oxirene-2,7-dione type: ligand / ID: 8 / Number of copies: 1 / Formula: A1AT1 |
|---|---|
| Molecular weight | Theoretical: 460.648 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOCONTINUUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




































