[English] 日本語
 Yorodumi
Yorodumi- EMDB-6559: Cryo-electron microscopy structure of ribosome-bound initiation f... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6559 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-electron microscopy structure of ribosome-bound initiation factor 2 70S IC II state | |||||||||
 Map data Map data | Reconstruction of the 70S-fMet-tRNAiMet-IF2-GDPNP complex, IC II state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | translation initiation / ribosome / initiation / Initiation Factor 2 / 70S initiation complex / prokaryotes / GTPase / translation GTPAse | |||||||||
| Function / homology |  Function and homology information Function and homology informationguanosine tetraphosphate binding / stringent response / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / ribosomal small subunit binding / misfolded RNA binding / Group I intron splicing / RNA folding / translational termination / transcriptional attenuation ...guanosine tetraphosphate binding / stringent response / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / ribosomal small subunit binding / misfolded RNA binding / Group I intron splicing / RNA folding / translational termination / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / four-way junction DNA binding / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation initiation factor activity / regulation of mRNA stability / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / response to cold / mRNA regulatory element binding translation repressor activity / regulation of DNA-templated transcription elongation / positive regulation of RNA splicing / transcription elongation factor complex / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / transcription antitermination / DNA endonuclease activity / molecular condensate scaffold activity / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / maintenance of translational fidelity / mRNA 5'-UTR binding / regulation of translation / large ribosomal subunit / transferase activity / ribosomal small subunit assembly / ribosome biogenesis / protein folding / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / hydrolase activity / mRNA binding / GTPase activity / GTP binding / DNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Sprink T / Ramrath DJF / Yamamoto H / Yamamoto K / Loerke J / Ismer J / Hildebrand PW / Scheerer P / Buerger J / Mielke T / Spahn CMT | |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2016 Journal: Sci Adv / Year: 2016Title: Structures of ribosome-bound initiation factor 2 reveal the mechanism of subunit association. Authors: Thiemo Sprink / David J F Ramrath / Hiroshi Yamamoto / Kaori Yamamoto / Justus Loerke / Jochen Ismer / Peter W Hildebrand / Patrick Scheerer / Jörg Bürger / Thorsten Mielke / Christian M T Spahn /  Abstract: Throughout the four phases of protein biosynthesis-initiation, elongation, termination, and recycling-the ribosome is controlled and regulated by at least one specified translational guanosine ...Throughout the four phases of protein biosynthesis-initiation, elongation, termination, and recycling-the ribosome is controlled and regulated by at least one specified translational guanosine triphosphatase (trGTPase). Although the structural basis for trGTPase interaction with the ribosome has been solved for the last three steps of translation, the high-resolution structure for the key initiation trGTPase, initiation factor 2 (IF2), complexed with the ribosome, remains elusive. We determine the structure of IF2 complexed with a nonhydrolyzable guanosine triphosphate analog and initiator fMet-tRNAi (Met) in the context of the Escherichia coli ribosome to 3.7-Å resolution using cryo-electron microscopy. The structural analysis reveals previously unseen intrinsic conformational modes of the 70S initiation complex, establishing the mutual interplay of IF2 and initator transfer RNA (tRNA) with the ribsosome and providing the structural foundation for a mechanistic understanding of the final steps of translation initiation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6559.map.gz emd_6559.map.gz | 163.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6559-v30.xml emd-6559-v30.xml emd-6559.xml emd-6559.xml | 13.1 KB 13.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6559.png emd_6559.png | 347 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6559 http://ftp.pdbj.org/pub/emdb/structures/EMD-6559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6559 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6559 | HTTPS FTP |
-Related structure data
| Related structure data |  3jcjMC  3285C  3jcnC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6559.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6559.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of the 70S-fMet-tRNAiMet-IF2-GDPNP complex, IC II state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.025 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
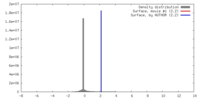
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Escherichia coli Initiation Factor 2 stalled on Escherichia coli ...
| Entire | Name: Escherichia coli Initiation Factor 2 stalled on Escherichia coli 70S ribosomes by the non-hydrolysable GTP analogue GDPNP in the presence of fMet-tRNAiMet and mRNA |
|---|---|
| Components |
|
-Supramolecule #1000: Escherichia coli Initiation Factor 2 stalled on Escherichia coli ...
| Supramolecule | Name: Escherichia coli Initiation Factor 2 stalled on Escherichia coli 70S ribosomes by the non-hydrolysable GTP analogue GDPNP in the presence of fMet-tRNAiMet and mRNA type: sample / ID: 1000 / Number unique components: 3 |
|---|---|
| Molecular weight | Experimental: 2.5 MDa / Theoretical: 2.6 MDa |
-Supramolecule #1: 70S ribosome
| Supramolecule | Name: 70S ribosome / type: complex / ID: 1 / Name.synonym: 70S / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 2.5 MDa / Theoretical: 2.6 MDa |
-Macromolecule #1: fMet-tRNAiMet
| Macromolecule | Name: fMet-tRNAiMet / type: rna / ID: 1 / Name.synonym: initiator tRNA / Classification: TRANSFER / Structure: DOUBLE HELIX / Synthetic?: No |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #2: Initiation Factor 2
| Macromolecule | Name: Initiation Factor 2 / type: protein_or_peptide / ID: 2 / Name.synonym: IF2 / Details: bound to GNPPNP / Number of copies: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 97 KDa / Theoretical: 97 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Translation initiation factor IF-2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 20 mM HEPES-KOH, pH 7.5, 15 mM magnesium acetate, 150 mM potassium acetate, 4 mM 2-mercapthoethanol, 2 mM spermidine, 0.05 mM spermine |
|---|---|
| Grid | Details: Quantifoil R3-3 Cu 300 mesh with 2 nm carbon support film |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK I / Method: Blot for 2-4 seconds before plunging. |
- Electron microscopy #1
Electron microscopy #1
| Microscopy ID | 1 |
|---|---|
| Microscope | FEI POLARA 300 |
| Date | Aug 26, 2013 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 918 / Average electron dose: 20 e/Å2 / Details: Automated data collection using Leginon |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 39000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 7.18 µm / Nominal defocus min: 0.64 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Electron microscopy #2
Electron microscopy #2
| Microscopy ID | 2 |
|---|---|
| Microscope | FEI POLARA 300 |
| Date | Jun 10, 2015 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 2797 / Average electron dose: 20 e/Å2 / Details: Automated data collection using Leginon |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 39000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 7.57 µm / Nominal defocus min: 0.19 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | To avoid overfitting, the data was refined in a resolution-limited scheme using SPIDER. |
|---|---|
| CTF correction | Details: CTFFIND4 |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.7 Å / Resolution method: OTHER / Software - Name: EMAN2, CTFFIND4, SPIDER, SPARX Details: Final maps were calculated from two combined datasets. To avoid overfitting, the data were refined in a resolution-limited scheme using SPIDER. Number images used: 54585 |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)