[English] 日本語
 Yorodumi
Yorodumi- EMDB-64903: Cryo-EM structure of formate dehydrogenase from Shewanella oneide... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of formate dehydrogenase from Shewanella oneidensis MR-1 (SoFdhAB) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Formate dehydrogenase / carbon dioxide reduction / tungsten (W)-dependent / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationformate dehydrogenase-N / formate dehydrogenase (quinone) activity / formate oxidation / formate dehydrogenase complex / molybdopterin cofactor binding / cell envelope / anaerobic respiration / 4 iron, 4 sulfur cluster binding / electron transfer activity / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.75 Å | |||||||||
 Authors Authors | Liu W / Zhang L | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2026 Journal: Nat Commun / Year: 2026Title: An interfacial-intramolecular electron highway for accelerated electrocatalytic CO reduction by an O-tolerant formate dehydrogenase. Authors: Weisong Liu / Peng Zhang / Xiufeng Wang / Kuncheng Zhang / Wenhua Yang / Huijuan Cui / Jun Liu / Junsong Sun / Chun You / Haiyang Cui / Zhiguang Zhu / Lingling Zhang /  Abstract: Bioelectrocatalytic CO reduction offers a sustainable route for CO bioconversion, yet remains limited by interfacial-intramolecular electron transfer and oxygen sensitivity. Here, we mine a formate ...Bioelectrocatalytic CO reduction offers a sustainable route for CO bioconversion, yet remains limited by interfacial-intramolecular electron transfer and oxygen sensitivity. Here, we mine a formate dehydrogenase from Shewanella oneidensis MR-1 (SoFdhAB) featuring completely oxygen tolerant and direct-electron-transfer (DET) electrocatalytic performances. Cryo-electron microscopy (Cryo-EM) analysis reveals an intramolecular electron highway comprising five [4Fe-4S] clusters, a regional face-face contact facilitating interfacial ET, and a unique oxygen resistance mechanism different from inactivation-activation. By acquiring a beneficial variant SoFdhAB-Y94S, a direct bioelectrocatalytic CO reduction system is constructed, accumulating 2.88 ± 0.03 mmol formate in 64 hours with a steady rate of 45.3 ± 0.5 μmol h cm and a Faradaic efficiency of 93.1 ± 5.2%. The merits of oxygen tolerance and efficient (electro)catalytic property endow SoFdhAB a robust enzyme adopted in potential application scenarios, and the inherent DET capability may inspire the interfacial engineering of other oxidoreductases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_64903.map.gz emd_64903.map.gz | 32.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-64903-v30.xml emd-64903-v30.xml emd-64903.xml emd-64903.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_64903.png emd_64903.png | 99.8 KB | ||
| Filedesc metadata |  emd-64903.cif.gz emd-64903.cif.gz | 7.1 KB | ||
| Others |  emd_64903_additional_1.map.gz emd_64903_additional_1.map.gz emd_64903_half_map_1.map.gz emd_64903_half_map_1.map.gz emd_64903_half_map_2.map.gz emd_64903_half_map_2.map.gz | 32.9 MB 59.5 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-64903 http://ftp.pdbj.org/pub/emdb/structures/EMD-64903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-64903 | HTTPS FTP |
-Related structure data
| Related structure data |  9vapMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_64903.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_64903.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_64903_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_64903_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_64903_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Heterodimer composed of formate dehydrogenase molybdopterin-bindi...
| Entire | Name: Heterodimer composed of formate dehydrogenase molybdopterin-binding subunit FdhA and 4Fe-4S dicluster domain-containing protein |
|---|---|
| Components |
|
-Supramolecule #1: Heterodimer composed of formate dehydrogenase molybdopterin-bindi...
| Supramolecule | Name: Heterodimer composed of formate dehydrogenase molybdopterin-binding subunit FdhA and 4Fe-4S dicluster domain-containing protein type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) |
| Molecular weight | Theoretical: 122 KDa |
-Macromolecule #1: Formate dehydrogenase molybdopterin-binding subunit FdhA
| Macromolecule | Name: Formate dehydrogenase molybdopterin-binding subunit FdhA type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: formate dehydrogenase-N |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) |
| Molecular weight | Theoretical: 105.996156 KDa |
| Recombinant expression | Organism:  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) |
| Sequence | String: MRLTRKTDPV VEPKVPALGL NRRQFLKSAG LATGGIAAAS MLGTGMMRKA EAKDIPHNAP TEVKRTICSH CSVGCGVYAE VQNGVWTGQ EPAFDHPFNQ GGHCAKGAAL REHGHGEKRL KYPMKLEGGK WKKISWDQAI NEVGDKMMAI RQESGPDSIY F MGSAKFSN ...String: MRLTRKTDPV VEPKVPALGL NRRQFLKSAG LATGGIAAAS MLGTGMMRKA EAKDIPHNAP TEVKRTICSH CSVGCGVYAE VQNGVWTGQ EPAFDHPFNQ GGHCAKGAAL REHGHGEKRL KYPMKLEGGK WKKISWDQAI NEVGDKMMAI RQESGPDSIY F MGSAKFSN EQAYLYRKFA ALWGTNNVDH SARICHSTTV AGVANTWGYG AQTNSVNDIR HSKCILFVGS NPSEAHPVAM QH ILVAKER GAKIIVVDPR FTRTAAKSDE YVHIRPGTDI PFIYGLLWHI FENGWEDKDF IKRRVYGMER IREEVKKYTP EEV ENVVGA PKAQMYRVAK MMAETKPGSI VWCMGGTQHH VGNANTRSYC ILQLALGNMG VTGGGTNIFR GHDNVQGASD FGLS FDDLP GYFGLTSGSW AHWANVWDLD PKWVTSRFDQ GEYLGQSPQT SPGIPCSRWH DGVLEDKTKI AQKDNIRLAF FWGQS VNTE TRGREVRQAL DKMDTVVVVD PFPTMAGVMH QRKDGVYLLP AATQFETYGS VSATNRSIQW RSKVIEPLFE SLPDHV IMC KLAKKVGIDK ELFKHIKVNG EEPLIEDIVR EYNRGMWTIG YTGQSPERLK MHQENWGTFN VDSLEAPGGP AKGETYG LP WPCWGTPEMK HPGSHILYNE TKHVKDGGGS FRARFGVERN GVNLLSEEAY SAGSEIQDGY PEFTADMLKQ LGWWDDLT E DEKKYAEGKN WKTDISGGIQ RVVIKHGCIP YGNGKARAVV WNFPDDIPLH REPLYTPRRD LVAKYPTYED RMVARLPTL YKSIQDKDFA KDFPLALTSG RLVEYEGGGE ETRSNPWLAE LQQEMFIEIS PADAADRGIR DGDNVFVHSP EGAKITVKAM VTPRVVPGE CFMPYHFAGV FEGESLAKNY PEGTVPYVIG ESANTILTYG YDVVTQMQET KSSLCQISKA UniProtKB: Formate dehydrogenase molybdopterin-binding subunit FdhA |
-Macromolecule #2: 4Fe-4S dicluster domain-containing protein
| Macromolecule | Name: 4Fe-4S dicluster domain-containing protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) |
| Molecular weight | Theoretical: 22.411707 KDa |
| Recombinant expression | Organism:  Shewanella oneidensis MR-1 (bacteria) Shewanella oneidensis MR-1 (bacteria) |
| Sequence | String: MAVMKFLCDT KRCIECNGCV TACKNENDSA LEWGIQRRRV VTINDGQPGE ASISVACMHC TDAPCMAVCP ADCFYRTDDG IVLHNKDTC IGCGYCFYAC PFGAPQFPKK TAFGSRGKMD KCTFCAGGPE ETFSEAEHKK YGANRIAEGK LPMCAELCAT K ALLAGDAE ...String: MAVMKFLCDT KRCIECNGCV TACKNENDSA LEWGIQRRRV VTINDGQPGE ASISVACMHC TDAPCMAVCP ADCFYRTDDG IVLHNKDTC IGCGYCFYAC PFGAPQFPKK TAFGSRGKMD KCTFCAGGPE ETFSEAEHKK YGANRIAEGK LPMCAELCAT K ALLAGDAE VVSNIYRQRM ASRGNPNVIW GYNPKTGEIN HHHHHH |
-Macromolecule #3: 2-AMINO-5,6-DIMERCAPTO-7-METHYL-3,7,8A,9-TETRAHYDRO-8-OXA-1,3,9,1...
| Macromolecule | Name: 2-AMINO-5,6-DIMERCAPTO-7-METHYL-3,7,8A,9-TETRAHYDRO-8-OXA-1,3,9,10-TETRAAZA-ANTHRACEN-4-ONE GUANOSINE DINUCLEOTIDE type: ligand / ID: 3 / Number of copies: 2 / Formula: MGD |
|---|---|
| Molecular weight | Theoretical: 740.557 Da |
| Chemical component information | 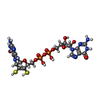 ChemComp-MGD: |
-Macromolecule #4: IRON/SULFUR CLUSTER
| Macromolecule | Name: IRON/SULFUR CLUSTER / type: ligand / ID: 4 / Number of copies: 5 / Formula: SF4 |
|---|---|
| Molecular weight | Theoretical: 351.64 Da |
| Chemical component information |  ChemComp-FS1: |
-Macromolecule #5: HYDROSULFURIC ACID
| Macromolecule | Name: HYDROSULFURIC ACID / type: ligand / ID: 5 / Number of copies: 1 / Formula: H2S |
|---|---|
| Molecular weight | Theoretical: 34.081 Da |
| Chemical component information |  ChemComp-H2S: |
-Macromolecule #6: TUNGSTEN ION
| Macromolecule | Name: TUNGSTEN ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: W |
|---|---|
| Molecular weight | Theoretical: 183.84 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 0.1 M Tris-HCl, 0.1 M NaCl |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)












































