[English] 日本語
 Yorodumi
Yorodumi- EMDB-44496: Cryo-EM co-structure of AcrB with the CU032 efflux pump inhibitor -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM co-structure of AcrB with the CU032 efflux pump inhibitor | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AcrB Multidrug Efflux Pump / TRANSLOCASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationxenobiotic detoxification by transmembrane export across the cell outer membrane / efflux pump complex / periplasmic side of plasma membrane / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / outer membrane-bounded periplasmic space / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
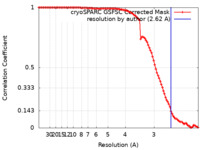
| Method | single particle reconstruction / cryo EM / Resolution: 2.62 Å | |||||||||
 Authors Authors | Su CC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: mBio / Year: 2023 Journal: mBio / Year: 2023Title: Bacterial efflux pump modulators prevent bacterial growth in macrophages and under broth conditions that mimic the host environment. Authors: Samual C Allgood / Chih-Chia Su / Amy L Crooks / Christian T Meyer / Bojun Zhou / Meredith D Betterton / Michael R Barbachyn / Edward W Yu / Corrella S Detweiler /  Abstract: Bacterial efflux pumps are critical for resistance to antibiotics and for virulence. We previously identified small molecules that inhibit efflux pumps (efflux pump modulators, EPMs) and prevent ...Bacterial efflux pumps are critical for resistance to antibiotics and for virulence. We previously identified small molecules that inhibit efflux pumps (efflux pump modulators, EPMs) and prevent pathogen replication in host cells. Here, we used medicinal chemistry to increase the activity of the EPMs against pathogens in cells into the nanomolar range. We show by cryo-electron microscopy that these EPMs bind an efflux pump subunit. In broth culture, the EPMs increase the potency (activity), but not the efficacy (maximum effect), of antibiotics. We also found that bacterial exposure to the EPMs appear to enable the accumulation of a toxic metabolite that would otherwise be exported by efflux pumps. Thus, inhibitors of bacterial efflux pumps could interfere with infection not only by potentiating antibiotics, but also by allowing toxic waste products to accumulate within bacteria, providing an explanation for why efflux pumps are needed for virulence in the absence of antibiotics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44496.map.gz emd_44496.map.gz | 97.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44496-v30.xml emd-44496-v30.xml emd-44496.xml emd-44496.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44496_fsc.xml emd_44496_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_44496.png emd_44496.png | 42.9 KB | ||
| Masks |  emd_44496_msk_1.map emd_44496_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44496.cif.gz emd-44496.cif.gz | 6.1 KB | ||
| Others |  emd_44496_half_map_1.map.gz emd_44496_half_map_1.map.gz emd_44496_half_map_2.map.gz emd_44496_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44496 http://ftp.pdbj.org/pub/emdb/structures/EMD-44496 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44496 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44496 | HTTPS FTP |
-Validation report
| Summary document |  emd_44496_validation.pdf.gz emd_44496_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_44496_full_validation.pdf.gz emd_44496_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_44496_validation.xml.gz emd_44496_validation.xml.gz | 18.3 KB | Display | |
| Data in CIF |  emd_44496_validation.cif.gz emd_44496_validation.cif.gz | 23.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44496 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44496 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44496 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-44496 | HTTPS FTP |
-Related structure data
| Related structure data |  9bfhMC  9bfmC  9bfnC  9bftC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44496.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44496.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44496_msk_1.map emd_44496_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
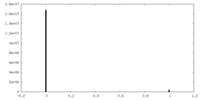
| Density Histograms |
-Half map: #2
| File | emd_44496_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
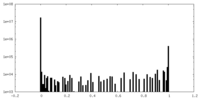
| Density Histograms |
-Half map: #1
| File | emd_44496_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
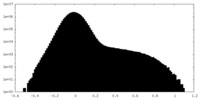
| Density Histograms |
- Sample components
Sample components
-Entire : H6PD
| Entire | Name: H6PD |
|---|---|
| Components |
|
-Supramolecule #1: H6PD
| Supramolecule | Name: H6PD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Multidrug efflux pump subunit AcrB
| Macromolecule | Name: Multidrug efflux pump subunit AcrB / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 113.66518 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR ...String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR TSGVGDVQLF GSQYAMRIWM NPNELNKFQL TPVDVITAIK AQNAQVAAGQ LGGTPPVKGQ QLNASIIAQT RL TSTEEFG KILLKVNQDG SRVLLRDVAK IELGGENYDI IAEFNGQPAS GLGIKLATGA NALDTAAAIR AELAKMEPFF PSG LKIVYP YDTTPFVKIS IHEVVKTLVE AIILVFLVMY LFLQNFRATL IPTIAVPVVL LGTFAVLAAF GFSINTLTMF GMVL AIGLL VDDAIVVVEN VERVMAEEGL PPKEATRKSM GQIQGALVGI AMVLSAVFVP MAFFGGSTGA IYRQFSITIV SAMAL SVLV ALILTPALCA TMLKPIAKGD HGEGKKGFFG WFNRMFEKST HHYTDSVGGI LRSTGRYLVL YLIIVVGMAY LFVRLP SSF LPDEDQGVFM TMVQLPAGAT QERTQKVLNE VTHYYLTKEK NNVESVFAVN GFGFAGRGQN TGIAFVSLKD WADRPGE EN KVEAITMRAT RAFSQIKDAM VFAFNLPAIV ELGTATGFDF ELIDQAGLGH EKLTQARNQL LAEAAKHPDM LTSVRPNG L EDTPQFKIDI DQEKAQALGV SINDINTTLG AAWGGSYVND FIDRGRVKKV YVMSEAKYRM LPDDIGDWYV RAADGQMVP FSAFSSSRWE YGSPRLERYN GLPSMEILGQ AAPGKSTGEA MELMEQLASK LPTGVGYDWT GMSYQERLSG NQAPSLYAIS LIVVFLCLA ALYESWSIPF SVMLVVPLGV IGALLAATFR GLTNDVYFQV GLLTTIGLSA KNAILIVEFA KDLMDKEGKG L IEATLDAV RMRLRPILMT SLAFILGVMP LVISTGAGSG AQNAVGTGVM GGMVTATVLA IFFVPVFFVV VRRRFSRKNE DI EHSHTVD HH UniProtKB: Multidrug efflux pump subunit AcrB |
-Macromolecule #2: (2S)-1-[(3R)-3-aminopyrrolidin-1-yl]-3-(3,4-dichlorophenoxy)propa...
| Macromolecule | Name: (2S)-1-[(3R)-3-aminopyrrolidin-1-yl]-3-(3,4-dichlorophenoxy)propan-2-ol type: ligand / ID: 2 / Number of copies: 1 / Formula: A1AN8 |
|---|---|
| Molecular weight | Theoretical: 305.2 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 5 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | This is from a heterogeneous and impure protein sample. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 29.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-9bfh: |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X