+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of the human cardiac F-actin mutant A331P | |||||||||
 Map data Map data | Final map used for model building. Postprocessed using phenix density modification. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | actin / cardiac / human / sarcomere / CONTRACTILE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic actin-based contraction involved in cell motility / actin-myosin filament sliding / cardiac myofibril assembly / actin filament-based movement / Formation of the dystrophin-glycoprotein complex (DGC) / cardiac muscle tissue morphogenesis / Striated Muscle Contraction / actomyosin structure organization / Regulation of CDH1 Function / I band ...cytoplasmic actin-based contraction involved in cell motility / actin-myosin filament sliding / cardiac myofibril assembly / actin filament-based movement / Formation of the dystrophin-glycoprotein complex (DGC) / cardiac muscle tissue morphogenesis / Striated Muscle Contraction / actomyosin structure organization / Regulation of CDH1 Function / I band / RHOB GTPase cycle / microfilament motor activity / myosin binding / heart contraction / mesenchyme migration / skeletal muscle thin filament assembly / RHOA GTPase cycle / cardiac muscle contraction / actin filament organization / sarcomere / filopodium / actin filament / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / lamellipodium / actin cytoskeleton / cell body / blood microparticle / response to ethanol / response to xenobiotic stimulus / focal adhesion / hydrolase activity / positive regulation of gene expression / negative regulation of apoptotic process / glutamatergic synapse / : / extracellular exosome / ATP binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Doran MH / Sousa D / Rynkiewicz MJ / Lehman W / Cammarato A | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of human cardiac actin Authors: Doran MH / Rynkiewicz MJ / Sousa D / Cammarato A / Lehman W | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44153.map.gz emd_44153.map.gz | 6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44153-v30.xml emd-44153-v30.xml emd-44153.xml emd-44153.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_44153.png emd_44153.png | 99.1 KB | ||
| Masks |  emd_44153_msk_1.map emd_44153_msk_1.map | 83.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44153.cif.gz emd-44153.cif.gz | 6.5 KB | ||
| Others |  emd_44153_additional_1.map.gz emd_44153_additional_1.map.gz emd_44153_additional_2.map.gz emd_44153_additional_2.map.gz emd_44153_half_map_1.map.gz emd_44153_half_map_1.map.gz emd_44153_half_map_2.map.gz emd_44153_half_map_2.map.gz | 64.6 MB 78.1 MB 65.2 MB 65.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44153 http://ftp.pdbj.org/pub/emdb/structures/EMD-44153 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44153 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44153 | HTTPS FTP |
-Related structure data
| Related structure data |  9b3qMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44153.map.gz / Format: CCP4 / Size: 6.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44153.map.gz / Format: CCP4 / Size: 6.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map used for model building. Postprocessed using phenix density modification. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.058 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44153_msk_1.map emd_44153_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
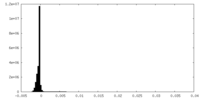
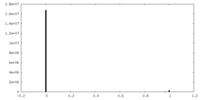
| Density Histograms |
-Additional map: Final refinement map before post-processing.
| File | emd_44153_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final refinement map before post-processing. | ||||||||||||
| Projections & Slices |
| ||||||||||||
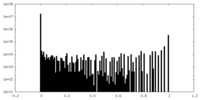
| Density Histograms |
-Additional map: Relion postprocessed map.
| File | emd_44153_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Relion postprocessed map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
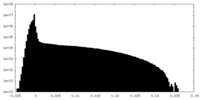
| Density Histograms |
-Half map: Half map 2
| File | emd_44153_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
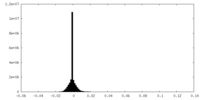
| Density Histograms |
-Half map: Half map 1
| File | emd_44153_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human cardiac F-actin with mutation A331P
| Entire | Name: Human cardiac F-actin with mutation A331P |
|---|---|
| Components |
|
-Supramolecule #1: Human cardiac F-actin with mutation A331P
| Supramolecule | Name: Human cardiac F-actin with mutation A331P / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: ACTC was expressed in Sf21 insect cells, using recombinant baculoviruses, and purified via gelsolin affinity chromatography |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 420 KDa |
-Macromolecule #1: Actin, alpha cardiac muscle 1
| Macromolecule | Name: Actin, alpha cardiac muscle 1 / type: protein_or_peptide / ID: 1 / Details: Human cardiac actin with the mutation A331P / Number of copies: 3 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.103945 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MCDDEETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIE(HIC)G IIT NWDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLD SG DGVTHNVPIY ...String: MCDDEETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIE(HIC)G IIT NWDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLD SG DGVTHNVPIY EGYALPHAIM RLDLAGRDLT DYLMKILTER GYSFVTTAER EIVRDIKEKL CYVALDFENE MATAASSS S LEKSYELPDG QVITIGNERF RCPETLFQPS FIGMESAGIH ETTYNSIMKC DIDIRKDLYA NNVLSGGTTM YPGIADRMQ KEITALAPST MKIKIIPPPE RKYSVWIGGS ILASLSTFQQ MWISKQEYDE AGPSIVHRKC F UniProtKB: Actin, alpha cardiac muscle 1 |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 3 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Details: 2 mmolL-1 Tris (pH 8), 0.2 mmolL-1 CaCl2, 0.2 mmolL-1 ATP, 0.5 mmolL-1 b-mercaptoethanol, 0.002% NaN3 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 8.0 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 27.93 Å Applied symmetry - Helical parameters - Δ&Phi: -166.45 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 140667 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: OTHER / Details: featureless cylinder |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller








 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)