+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a membrane transport protein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Transport / Membrane / Inhibitor / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationDefective SLC6A3 causes Parkinsonism-dystonia infantile (PKDYS) / Defective SLC6A3 causes Parkinsonism-dystonia infantile (PKDYS) / Dopamine clearance from the synaptic cleft / amine binding / adenohypophysis development / dopamine uptake / hyaloid vascular plexus regression / dopamine binding / norepinephrine:sodium symporter activity / dopamine:sodium symporter activity ...Defective SLC6A3 causes Parkinsonism-dystonia infantile (PKDYS) / Defective SLC6A3 causes Parkinsonism-dystonia infantile (PKDYS) / Dopamine clearance from the synaptic cleft / amine binding / adenohypophysis development / dopamine uptake / hyaloid vascular plexus regression / dopamine binding / norepinephrine:sodium symporter activity / dopamine:sodium symporter activity / norepinephrine transport / regulation of dopamine metabolic process / dopaminergic synapse / neurotransmitter transmembrane transporter activity / dopamine transport / flotillin complex / monoamine transmembrane transporter activity / Na+/Cl- dependent neurotransmitter transporters / monoamine transport / dopamine catabolic process / positive regulation of multicellular organism growth / neurotransmitter transport / response to iron ion / dopamine biosynthetic process / dopamine uptake involved in synaptic transmission / amino acid transport / heterocyclic compound binding / neuronal cell body membrane / sodium ion transmembrane transport / prepulse inhibition / axon terminus / response to cAMP / lactation / protein phosphatase 2A binding / response to cocaine / locomotory behavior / response to nicotine / cognition / sensory perception of smell / presynaptic membrane / protease binding / postsynaptic membrane / response to ethanol / neuron projection / response to xenobiotic stimulus / membrane raft / axon / signaling receptor binding / neuronal cell body / protein-containing complex binding / cell surface / membrane / metal ion binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
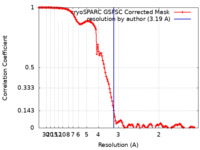
| Method | single particle reconstruction / cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Srivastava DK / Gouaux E | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Structure of the human dopamine transporter and mechanisms of inhibition Authors: Srivastava DK / Navratna V / Tosh DK / Chinn A / Sk MF / Tajkhorshid E / Jacobson KA / Gouaux E | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43128.map.gz emd_43128.map.gz | 203.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43128-v30.xml emd-43128-v30.xml emd-43128.xml emd-43128.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43128_fsc.xml emd_43128_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_43128.png emd_43128.png | 59.9 KB | ||
| Masks |  emd_43128_msk_1.map emd_43128_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-43128.cif.gz emd-43128.cif.gz | 6.8 KB | ||
| Others |  emd_43128_half_map_1.map.gz emd_43128_half_map_1.map.gz emd_43128_half_map_2.map.gz emd_43128_half_map_2.map.gz | 200.5 MB 200.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43128 http://ftp.pdbj.org/pub/emdb/structures/EMD-43128 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43128 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43128 | HTTPS FTP |
-Validation report
| Summary document |  emd_43128_validation.pdf.gz emd_43128_validation.pdf.gz | 972.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43128_full_validation.pdf.gz emd_43128_full_validation.pdf.gz | 972.4 KB | Display | |
| Data in XML |  emd_43128_validation.xml.gz emd_43128_validation.xml.gz | 21.5 KB | Display | |
| Data in CIF |  emd_43128_validation.cif.gz emd_43128_validation.cif.gz | 27.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43128 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43128 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43128 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43128 | HTTPS FTP |
-Related structure data
| Related structure data |  8vbyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43128.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43128.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.743 Å | ||||||||||||||||||||||||||||||||||||
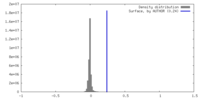
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_43128_msk_1.map emd_43128_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
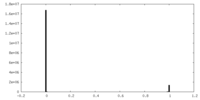
| Density Histograms |
-Half map: #2
| File | emd_43128_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
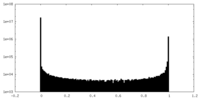
| Density Histograms |
-Half map: #1
| File | emd_43128_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
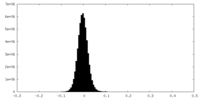
| Density Histograms |
- Sample components
Sample components
+Entire : Sodium dependent dopamine transporter in complex with inhibitor
+Supramolecule #1: Sodium dependent dopamine transporter in complex with inhibitor
+Macromolecule #1: Sodium-dependent dopamine transporter
+Macromolecule #2: methyl (1R,2S,3S,5S)-3-(4-fluorophenyl)-8-methyl-8-azabicyclo[3.2...
+Macromolecule #3: methyl (1S,2R,3S,4R,5R)-4-{2-[(5-chlorothiophen-2-yl)ethynyl]-6-(...
+Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #5: CHOLESTEROL HEMISUCCINATE
+Macromolecule #6: DECANE
+Macromolecule #7: PENTANE
+Macromolecule #8: HEPTANE
+Macromolecule #9: ZINC ION
+Macromolecule #10: SODIUM ION
+Macromolecule #11: DODECANE
+Macromolecule #12: PENTADECANE
+Macromolecule #13: N-OCTANE
+Macromolecule #14: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.0 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 288.15 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||
| Details | monodisperse sample |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 6 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 165000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)