+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Arp2/3 branch junction complex, ADP state | ||||||||||||
 Map data Map data | Cryosparc final map, sharpened | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | actin / arp2/3 / cytoskeleton / branch / CYTOSOLIC PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationRegulation of actin dynamics for phagocytic cup formation / RHO GTPases Activate WASPs and WAVEs / Clathrin-mediated endocytosis / actin cortical patch organization / medial cortex / Neutrophil degranulation / actin filament branching / cell cortex of cell tip / actin cortical patch assembly / Arp2/3 protein complex ...Regulation of actin dynamics for phagocytic cup formation / RHO GTPases Activate WASPs and WAVEs / Clathrin-mediated endocytosis / actin cortical patch organization / medial cortex / Neutrophil degranulation / actin filament branching / cell cortex of cell tip / actin cortical patch assembly / Arp2/3 protein complex / Arp2/3 complex-mediated actin nucleation / actin cortical patch / cell tip / regulation of actin filament polymerization / mating projection tip / cortical actin cytoskeleton organization / cytoskeletal motor activator activity / cell division site / myosin heavy chain binding / tropomyosin binding / actin filament bundle / troponin I binding / filamentous actin / mesenchyme migration / establishment or maintenance of cell polarity / cortical cytoskeleton / skeletal muscle myofibril / actin filament bundle assembly / striated muscle thin filament / skeletal muscle thin filament assembly / actin monomer binding / skeletal muscle fiber development / stress fiber / titin binding / actin filament polymerization / filopodium / actin filament / structural constituent of cytoskeleton / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / endocytosis / calcium-dependent protein binding / actin filament binding / lamellipodium / cell body / cell cortex / protein-macromolecule adaptor activity / protein domain specific binding / hydrolase activity / calcium ion binding / positive regulation of gene expression / magnesium ion binding / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |    | ||||||||||||
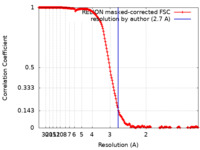
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | ||||||||||||
 Authors Authors | Chavali SS / Chou SZ / Sindelar CV | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Cryo-EM structures reveal how phosphate release from Arp3 weakens actin filament branches formed by Arp2/3 complex. Authors: Sai Shashank Chavali / Steven Z Chou / Wenxiang Cao / Thomas D Pollard / Enrique M De La Cruz / Charles V Sindelar /  Abstract: Arp2/3 complex nucleates branched actin filaments for cell and organelle movements. Here we report a 2.7 Å resolution cryo-EM structure of the mature branch junction formed by S. pombe Arp2/3 ...Arp2/3 complex nucleates branched actin filaments for cell and organelle movements. Here we report a 2.7 Å resolution cryo-EM structure of the mature branch junction formed by S. pombe Arp2/3 complex that provides details about interactions with both mother and daughter filaments. We determine a second structure at 3.2 Å resolution with the phosphate analog BeF bound with ADP to Arp3 and ATP bound to Arp2. In this ADP-BeF transition state the outer domain of Arp3 is rotated 2° toward the mother filament compared with the ADP state and makes slightly broader contacts with actin in both the mother and daughter filaments. Thus, dissociation of P from the ADP-P transition state reduces the interactions of Arp2/3 complex with the actin filaments and may contribute to the lower mechanical stability of mature branch junctions with ADP bound to the Arps. Our structures also reveal that the mother filament in contact with Arp2/3 complex is slightly bent and twisted, consistent with the preference of Arp2/3 complex binding curved actin filaments. The small degree of twisting constrains models of actin filament mechanics. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42787.map.gz emd_42787.map.gz | 483.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42787-v30.xml emd-42787-v30.xml emd-42787.xml emd-42787.xml | 31.4 KB 31.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42787_fsc.xml emd_42787_fsc.xml | 18.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_42787.png emd_42787.png | 148.4 KB | ||
| Filedesc metadata |  emd-42787.cif.gz emd-42787.cif.gz | 8.6 KB | ||
| Others |  emd_42787_additional_1.map.gz emd_42787_additional_1.map.gz emd_42787_half_map_1.map.gz emd_42787_half_map_1.map.gz emd_42787_half_map_2.map.gz emd_42787_half_map_2.map.gz | 255.6 MB 475.6 MB 475.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42787 http://ftp.pdbj.org/pub/emdb/structures/EMD-42787 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42787 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42787 | HTTPS FTP |
-Related structure data
| Related structure data |  8uxwMC  8uxxC  8uz0C  8uz1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42787.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42787.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryosparc final map, sharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.673 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Cryosparc final map, no sharpening
| File | emd_42787_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryosparc final map, no sharpening | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryosparc half map A
| File | emd_42787_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryosparc half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryosparc half map B
| File | emd_42787_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryosparc half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Arp2/3 branch complex (ADP state)
+Supramolecule #1: Arp2/3 branch complex (ADP state)
+Supramolecule #2: Arp2/3 complex
+Supramolecule #3: Actin
+Macromolecule #1: Actin-related protein 3
+Macromolecule #2: Actin-related protein 2
+Macromolecule #3: Actin-related protein 2/3 complex subunit 1
+Macromolecule #4: Actin-related protein 2/3 complex subunit 2
+Macromolecule #5: Actin-related protein 2/3 complex subunit 3
+Macromolecule #6: Actin-related protein 2/3 complex subunit 4
+Macromolecule #7: Actin-related protein 2/3 complex subunit 5
+Macromolecule #8: Actin, alpha skeletal muscle
+Macromolecule #9: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #10: MAGNESIUM ION
+Macromolecule #11: ADENOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Details: Grids were not glow discharged |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: The samples were incubated on the grid for 50 s and the extra solution was blotted using two Vitrobot filter papers (0.55/20 mm, Grade 595, Ted Pella) for 4 s at 0 blot force. The grids were ...Details: The samples were incubated on the grid for 50 s and the extra solution was blotted using two Vitrobot filter papers (0.55/20 mm, Grade 595, Ted Pella) for 4 s at 0 blot force. The grids were plunged into liquid ethane at ~180 degrees C with a wait time of 0.5 s.. |
| Details | Actin monomers with bound Ca2+ were converted to Mg2+-actin by equilibrating with 50 micromolar MgCl2 and 0.2 mM EGTA (pH 7.5) for 10 min on ice. Actin was polymerized in the presence of capping protein (CP) by sequentially mixing 8.75 micromolar Mg-ATP-actin monomers with 0.75 micromolar CP and equilibrated at room temperature for 1 h. In parallel, Arp2/3 complex (0.4 micromolar) in QB buffer was activated by mixing 0.85 micromolar GCN4-VCA and 50 micromolar ATP and incubated at 4 degrees for 1 h. The capped actin filaments sample was then gently mixed with an equal volume of activated Arp2/3 complex sample using cut pipette tips and equilibrated at 4 degrees for 5 min. Daughter filaments were formed and elongated in the presence of CP by adding 0.25 micromolar Mg-actin monomers, 50 micromolar ATP, and 40 nM CP and incubated for 5 min at room temperature. This step was subsequently repeated 4 more times, then aged for ~90 min before preparing grids for cryo-EM data collection. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV Details: Electron micrographs for image reconstructions were collected using Titan Krios equipped with X-cold field emission gun at 300 kV, Gatan image filter with slit width of 20 eV and a nanoprobe. |
| Details | The vitrified grids were screened for sample homogeneity and ice thickness in a Glacios 200 kV transmission electron microscope equipped with Gatan K2 summit camera. Electron micrographs for image reconstructions were collected using Titan Krios equipped with X-cold field emission gun at 300 kV, Gatan image filter with slit width of 20 eV and a nanoprobe. |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 8519 / Average exposure time: 3.3 sec. / Average electron dose: 51.4 e/Å2 Details: Each movie contains 41 frames with a frame time of 0.08 s. A dose rate of 28.4 counts/pixel/s and a physical pixel size of 1.346 Angstroms was used. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 64000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)