+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | DdmD monomer in complex with ssDNA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DdmD / helicase / nuclease / IMMUNE SYSTEM / IMMUNE SYSTEM-DNA complex | |||||||||
| Function / homology | Helicase/UvrB N-terminal domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.99 Å | |||||||||
 Authors Authors | Bravo JPK / Taylor DW | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Plasmid targeting and destruction by the DdmDE bacterial defence system. Authors: Jack P K Bravo / Delisa A Ramos / Rodrigo Fregoso Ocampo / Caiden Ingram / David W Taylor /   Abstract: Although eukaryotic Argonautes have a pivotal role in post-transcriptional gene regulation through nucleic acid cleavage, some short prokaryotic Argonaute variants (pAgos) rely on auxiliary nuclease ...Although eukaryotic Argonautes have a pivotal role in post-transcriptional gene regulation through nucleic acid cleavage, some short prokaryotic Argonaute variants (pAgos) rely on auxiliary nuclease factors for efficient foreign DNA degradation. Here we reveal the activation pathway of the DNA defence module DdmDE system, which rapidly eliminates small, multicopy plasmids from the Vibrio cholerae seventh pandemic strain (7PET). Through a combination of cryo-electron microscopy, biochemistry and in vivo plasmid clearance assays, we demonstrate that DdmE is a catalytically inactive, DNA-guided, DNA-targeting pAgo with a distinctive insertion domain. We observe that the helicase-nuclease DdmD transitions from an autoinhibited, dimeric complex to a monomeric state upon loading of single-stranded DNA targets. Furthermore, the complete structure of the DdmDE-guide-target handover complex provides a comprehensive view into how DNA recognition triggers processive plasmid destruction. Our work establishes a mechanistic foundation for how pAgos utilize ancillary factors to achieve plasmid clearance, and provides insights into anti-plasmid immunity in bacteria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41790.map.gz emd_41790.map.gz | 108.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41790-v30.xml emd-41790-v30.xml emd-41790.xml emd-41790.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41790.png emd_41790.png | 57.3 KB | ||
| Filedesc metadata |  emd-41790.cif.gz emd-41790.cif.gz | 6.1 KB | ||
| Others |  emd_41790_half_map_1.map.gz emd_41790_half_map_1.map.gz emd_41790_half_map_2.map.gz emd_41790_half_map_2.map.gz | 200.3 MB 200.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41790 http://ftp.pdbj.org/pub/emdb/structures/EMD-41790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41790 | HTTPS FTP |
-Validation report
| Summary document |  emd_41790_validation.pdf.gz emd_41790_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41790_full_validation.pdf.gz emd_41790_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_41790_validation.xml.gz emd_41790_validation.xml.gz | 15.6 KB | Display | |
| Data in CIF |  emd_41790_validation.cif.gz emd_41790_validation.cif.gz | 18.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41790 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41790 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41790 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41790 | HTTPS FTP |
-Related structure data
| Related structure data |  8u0wMC  8u0jC  8u0uC  8u3kC  9bqvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41790.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41790.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8332 Å | ||||||||||||||||||||||||||||||||||||
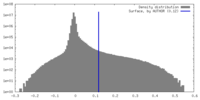
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_41790_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
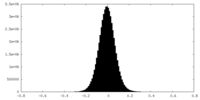
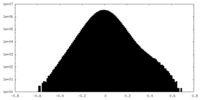
| Density Histograms |
-Half map: #2
| File | emd_41790_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
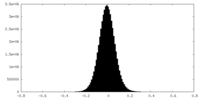
| Density Histograms |
- Sample components
Sample components
-Entire : DdmD
| Entire | Name: DdmD |
|---|---|
| Components |
|
-Supramolecule #1: DdmD
| Supramolecule | Name: DdmD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Helicase/UvrB N-terminal domain-containing protein
| Macromolecule | Name: Helicase/UvrB N-terminal domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 140.011953 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MFMYSVFTCR YWRYIMVRLI KHQQLLGEYC MNVSIEEFTH FDFQLVPEPS PLDLVITESL KNHIEVNGVK SGALLPLPFQ TGIGKTYTA LNFLLQQMLE QVRSELKEEN TGKKSKRLLY YVTDSVDNVV SAKADLLKLI EKQTVKGEPR FTLEQQEYLK A QIVHLPNQ ...String: MFMYSVFTCR YWRYIMVRLI KHQQLLGEYC MNVSIEEFTH FDFQLVPEPS PLDLVITESL KNHIEVNGVK SGALLPLPFQ TGIGKTYTA LNFLLQQMLE QVRSELKEEN TGKKSKRLLY YVTDSVDNVV SAKADLLKLI EKQTVKGEPR FTLEQQEYLK A QIVHLPNQ SEQLLQCSDA VLNDVLIGFN LNAERDVQAE WSAISGLRRH ASNPEVKISL NRQAGYFYRN LIDRLQKKQK GA DRVLLSG SLLASVETLL PGEKIRNGSA HVAFLTTSKF LKGFHNTRSR YSPLRDLSGA VLIIDEIDKQ NQVILSELCK QQA QDLIWA IRTLRANFRD HQLESSPRYD KIEDLFEPLR ERLEEFGTNW NLAFAFNTEG ANLNERPVRL FSDRSFTHVS SATH KLSLK SDFLRRKNLI FSDEKVEGSL IEKHGLLTRF VNEADVIYQW FLGTMRKAVF QYWENVRGLE IEVRENRSLE GTFQE AVQS LLTHFNLQEF ESAVYESFDT RGLRQSAGGK ANKLSSSKSY HHTGLKLVEV AHNQGTRDTV NCKASFLNTS PSGVLA DMV DAGAVILGIS ATARADTVIH NFDFKYLNER LGNKLLSLSR EQKQRVNNYY HSRRNYKDNG VVLTVKYLNS RDAFLDA LL EEYKPEARSS HFILNHYLGI AESEQAFVRS WLSKLLASIK AFISSPDNRY MLSLLNRTLD TTRQNINDFI QFCCDKWA K EFNVKTKTFF GVNADWMRLV GYDEISKHLN TELGKVVVFS TYASMGAGKN PDYAVNLALE GESLISVADV TYSTQLRSD IDSIYLEKPT QLLLSDDYSH TANQLCQFHQ ILSLQENGEL SPKSAENWCR QQLMGMSRER SLQQYHQTSD YQSAVRKYIE QAVGRAGRT SLKRKQILLF VDSGLKEILA EESRDPSLFS HEYVALVNKA KSAGKSIVED RAVRRLFNLA QRNNKDGMLS I KALVHRLH NQPASKSDIQ EWQDIRTQLL RYPTVAFQPE RFNRLYLQSM TKGYYRYQGN LDGDPNSFEF FDRVPYGDMV SE EDCSLAT LVQNQYVRPW FERKGFACSW QKEANVMTPI MFTNIYKGAL GEQAVEAVLT AFDFTFEEVP NSIYERFDNR VIF AGIEQP IWLDSKYWKH EGNESSEGYS SKIALVEEEF GPSKFIYVNA LGDTSKPIRY LNSCFVETSP QLAKVIEIPA LIDD SNADT NRTAVQELIK WLHHS UniProtKB: Helicase/UvrB N-terminal domain-containing protein |
-Macromolecule #2: DNA (5'-D(P*TP*TP*AP*TP*TP*TP*TP*TP*TP*TP*TP*T)-3')
| Macromolecule | Name: DNA (5'-D(P*TP*TP*AP*TP*TP*TP*TP*TP*TP*TP*TP*T)-3') / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 3.61437 KDa |
| Sequence | String: (DT)(DT)(DA)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.99 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 235731 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)