[English] 日本語
 Yorodumi
Yorodumi- EMDB-41322: Myxococcus xanthus EncA protein shell with compartmentalized SNAP... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Myxococcus xanthus EncA protein shell with compartmentalized SNAP-tag cargo protein | |||||||||
 Map data Map data | Myxococcus xanthus EncA protein shell with compartmentalized SNAP-tag cargo protein | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Encapsulin / nanocompartment / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationmethylated-DNA-[protein]-cysteine S-methyltransferase / methylated-DNA-[protein]-cysteine S-methyltransferase activity / encapsulin nanocompartment / iron ion transport / methylation / intracellular iron ion homeostasis / DNA repair / DNA binding / metal ion binding / nucleus Similarity search - Function | |||||||||
| Biological species |  Myxococcus xanthus DK 1622 (bacteria) / Myxococcus xanthus DK 1622 (bacteria) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.53 Å | |||||||||
 Authors Authors | Andreas MP / Kwon S / Giessen TW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2023 Journal: J Struct Biol / Year: 2023Title: Structure and heterogeneity of a highly cargo-loaded encapsulin shell. Authors: Seokmu Kwon / Michael P Andreas / Tobias W Giessen /  Abstract: Encapsulins are self-assembling protein nanocompartments able to selectively encapsulate dedicated cargo enzymes. Encapsulins are widespread across bacterial and archaeal phyla and are involved in ...Encapsulins are self-assembling protein nanocompartments able to selectively encapsulate dedicated cargo enzymes. Encapsulins are widespread across bacterial and archaeal phyla and are involved in oxidative stress resistance, iron storage, and sulfur metabolism. Encapsulin shells exhibit icosahedral geometry and consist of 60, 180, or 240 identical protein subunits. Cargo encapsulation is mediated by the specific interaction of targeting peptides or domains, found in all cargo proteins, with the interior surface of the encapsulin shell during shell self-assembly. Here, we report the 2.53 Å cryo-EM structure of a heterologously produced and highly cargo-loaded T3 encapsulin shell from Myxococcus xanthus and explore the systems' structural heterogeneity. We find that exceedingly high cargo loading results in the formation of substantial amounts of distorted and aberrant shells, likely caused by a combination of unfavorable steric clashes of cargo proteins and shell conformational changes. Based on our cryo-EM structure, we determine and analyze the targeting peptide-shell binding mode. We find that both ionic and hydrophobic interactions mediate targeting peptide binding. Our results will guide future attempts at rationally engineering encapsulins for biomedical and biotechnological applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41322.map.gz emd_41322.map.gz | 307.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41322-v30.xml emd-41322-v30.xml emd-41322.xml emd-41322.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
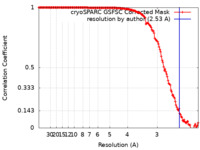
| FSC (resolution estimation) |  emd_41322_fsc.xml emd_41322_fsc.xml | 14.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_41322.png emd_41322.png | 184.9 KB | ||
| Others |  emd_41322_half_map_1.map.gz emd_41322_half_map_1.map.gz emd_41322_half_map_2.map.gz emd_41322_half_map_2.map.gz | 301.2 MB 301.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41322 http://ftp.pdbj.org/pub/emdb/structures/EMD-41322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41322 | HTTPS FTP |
-Related structure data
| Related structure data |  8tk7MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41322.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41322.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Myxococcus xanthus EncA protein shell with compartmentalized SNAP-tag cargo protein | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.10345 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_41322_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_41322_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Myxococcus xanthus EncA protein shell with compartmentalized SNAP...
| Entire | Name: Myxococcus xanthus EncA protein shell with compartmentalized SNAP-tag cargo protein |
|---|---|
| Components |
|
-Supramolecule #1: Myxococcus xanthus EncA protein shell with compartmentalized SNAP...
| Supramolecule | Name: Myxococcus xanthus EncA protein shell with compartmentalized SNAP-tag cargo protein type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Myxococcus xanthus DK 1622 (bacteria) Myxococcus xanthus DK 1622 (bacteria) |
-Supramolecule #2: Myxococcus xanthus EncA encapsulin shell protein
| Supramolecule | Name: Myxococcus xanthus EncA encapsulin shell protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 Details: Recombinantly expressed Myxococcus xanthus EncA encapsulin shell with T=3 icosahedral symmetry |
|---|---|
| Source (natural) | Organism:  Myxococcus xanthus DK 1622 (bacteria) Myxococcus xanthus DK 1622 (bacteria) |
-Supramolecule #3: SNAP-Tag/EncC targeting peptide
| Supramolecule | Name: SNAP-Tag/EncC targeting peptide / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 Details: Internalized SNAP-tag/EncC targeting peptide cargo protein |
|---|---|
| Source (natural) | Organism:  Myxococcus xanthus DK 1622 (bacteria) Myxococcus xanthus DK 1622 (bacteria) |
-Macromolecule #1: Type 1 encapsulin shell protein EncA
| Macromolecule | Name: Type 1 encapsulin shell protein EncA / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Myxococcus xanthus DK 1622 (bacteria) Myxococcus xanthus DK 1622 (bacteria) |
| Molecular weight | Theoretical: 31.691977 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPDFLGHAEN PLREEEWARL NETVIQVARR SLVGRRILDI YGPLGAGVQT VPYDEFQGVS PGAVDIVGEQ ETAMVFTDAR KFKTIPIIY KDFLLHWRDI EAARTHNMPL DVSAAAGAAA LCAQQEDELI FYGDARLGYE GLMTANGRLT VPLGDWTSPG G GFQAIVEA ...String: MPDFLGHAEN PLREEEWARL NETVIQVARR SLVGRRILDI YGPLGAGVQT VPYDEFQGVS PGAVDIVGEQ ETAMVFTDAR KFKTIPIIY KDFLLHWRDI EAARTHNMPL DVSAAAGAAA LCAQQEDELI FYGDARLGYE GLMTANGRLT VPLGDWTSPG G GFQAIVEA TRKLNEQGHF GPYAVVLSPR LYSQLHRIYE KTGVLEIETI RQLASDGVYQ SNRLRGESGV VVSTGRENMD LA VSMDMVA AYLGASRMNH PFRVLEALLL RIKHPDAICT LEGAGATERR UniProtKB: Type 1 encapsulin shell protein EncA |
-Macromolecule #2: Methylated-DNA--protein-cysteine methyltransferase
| Macromolecule | Name: Methylated-DNA--protein-cysteine methyltransferase / type: protein_or_peptide / ID: 2 Details: Amino acids 1-183 are unresolved SNAP-TAG. Amino acids 184-191 are unresolved linker. Amino acids 192-203 are EncC targeting peptide. Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.4826 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGPGSDKDCE MKRTTLDSPL GKLELSGCEQ GLHEIIFLGK GTSAADAVEV PAPAAVLGGP EPLMQATAWL NAYFHQPEAI EEFPVPALH HPVFQQESFT RQVLWKLLKV VKFGEVISYS HLAALAGNPA ATAAVKTALS GNPVPILIPC HRVVQGDLDV G GYEGGLAV ...String: MGPGSDKDCE MKRTTLDSPL GKLELSGCEQ GLHEIIFLGK GTSAADAVEV PAPAAVLGGP EPLMQATAWL NAYFHQPEAI EEFPVPALH HPVFQQESFT RQVLWKLLKV VKFGEVISYS HLAALAGNPA ATAAVKTALS GNPVPILIPC HRVVQGDLDV G GYEGGLAV KEWLLAHEGH RLGKRGGGSG GGSPEKRLTV GSLRR UniProtKB: Methylated-DNA--protein-cysteine methyltransferase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.4 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 150 mM NaCl, 20 mM Tris pH 7.5 | |||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 4092 pixel / Digitization - Dimensions - Height: 5760 pixel / Number grids imaged: 1 / Number real images: 2610 / Average exposure time: 3.2 sec. / Average electron dose: 49.26 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | The model was initially fit into the map using UCSF ChimeraX. It was then manually refined using Coot, followed by real-space refinement against the map using Phenix. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 89.9 / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-8tk7: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)