[English] 日本語
 Yorodumi
Yorodumi- EMDB-41078: Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | encapsulin / virus like particle / protein nanocompartment | |||||||||
| Function / homology | :  Function and homology information Function and homology information | |||||||||
| Biological species |  Acinetobacter baumannii 118362 (bacteria) Acinetobacter baumannii 118362 (bacteria) | |||||||||
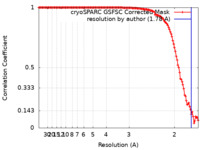
| Method | single particle reconstruction / cryo EM / Resolution: 1.78 Å | |||||||||
 Authors Authors | Andreas MP / Benisch R / Giessen TW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2024 Journal: Sci Adv / Year: 2024Title: A widespread bacterial protein compartment sequesters and stores elemental sulfur. Authors: Robert Benisch / Michael P Andreas / Tobias W Giessen /  Abstract: Subcellular compartments often serve to store nutrients or sequester labile or toxic compounds. As bacteria mostly do not possess membrane-bound organelles, they often have to rely on protein-based ...Subcellular compartments often serve to store nutrients or sequester labile or toxic compounds. As bacteria mostly do not possess membrane-bound organelles, they often have to rely on protein-based compartments. Encapsulins are one of the most prevalent protein-based compartmentalization strategies found in prokaryotes. Here, we show that desulfurase encapsulins can sequester and store large amounts of crystalline elemental sulfur. We determine the 1.78-angstrom cryo-EM structure of a 24-nanometer desulfurase-loaded encapsulin. Elemental sulfur crystals can be formed inside the encapsulin shell in a desulfurase-dependent manner with l-cysteine as the sulfur donor. Sulfur accumulation can be influenced by the concentration and type of sulfur source in growth medium. The selectively permeable protein shell allows the storage of redox-labile elemental sulfur by excluding cellular reducing agents, while encapsulation substantially improves desulfurase activity and stability. These findings represent an example of a protein compartment able to accumulate and store elemental sulfur. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41078.map.gz emd_41078.map.gz | 204.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41078-v30.xml emd-41078-v30.xml emd-41078.xml emd-41078.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41078_fsc.xml emd_41078_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_41078.png emd_41078.png | 174.4 KB | ||
| Filedesc metadata |  emd-41078.cif.gz emd-41078.cif.gz | 6.2 KB | ||
| Others |  emd_41078_half_map_1.map.gz emd_41078_half_map_1.map.gz emd_41078_half_map_2.map.gz emd_41078_half_map_2.map.gz | 198.3 MB 198.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41078 http://ftp.pdbj.org/pub/emdb/structures/EMD-41078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41078 | HTTPS FTP |
-Validation report
| Summary document |  emd_41078_validation.pdf.gz emd_41078_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41078_full_validation.pdf.gz emd_41078_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_41078_validation.xml.gz emd_41078_validation.xml.gz | 21.6 KB | Display | |
| Data in CIF |  emd_41078_validation.cif.gz emd_41078_validation.cif.gz | 27.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41078 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41078 | HTTPS FTP |
-Related structure data
| Related structure data |  8t6rMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41078.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41078.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_41078_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_41078_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell
| Entire | Name: Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell |
|---|---|
| Components |
|
-Supramolecule #1: Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell
| Supramolecule | Name: Acinetobacter baumannii 118362 family 2A cargo-loaded encapsulin shell type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Acinetinobacter baumannii 118362 family 2A encapsulin shell with internal cysteine desulfurase cargo protein |
|---|---|
| Source (natural) | Organism:  Acinetobacter baumannii 118362 (bacteria) Acinetobacter baumannii 118362 (bacteria) |
| Molecular weight | Theoretical: 1.88 MDa |
-Macromolecule #1: Major membrane protein I
| Macromolecule | Name: Major membrane protein I / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acinetobacter baumannii 118362 (bacteria) Acinetobacter baumannii 118362 (bacteria) |
| Molecular weight | Theoretical: 34.609195 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAKNTDKAQL ALGDHAARQL ANATKTAPQL STITPRWLTH LLQWIPVEAG IYRLNRVNNT DDIQVACTQR DEATLPQTFV DYAPEPREY FLNGVSTVLD VHTRVADLYS SPHDQIKEQL RLTIETIKER QESELINNPE YGLLASVTDD QRISTLNGPP T PDDLDDLL ...String: MAKNTDKAQL ALGDHAARQL ANATKTAPQL STITPRWLTH LLQWIPVEAG IYRLNRVNNT DDIQVACTQR DEATLPQTFV DYAPEPREY FLNGVSTVLD VHTRVADLYS SPHDQIKEQL RLTIETIKER QESELINNPE YGLLASVTDD QRISTLNGPP T PDDLDDLL RKVWKEPGFF LAHPDAIAAF GRECTRRGVP PPTVSLFGSQ FITWRGIPLI PSNKIPVEDG KTKILLLRVG EK RQGIVGL FQPGLAGEQS PGLSVRFMGI NRNAIASYLI SLYCSLAVLT DDALAVLDDV EVDKYHDYPV NYK UniProtKB: UNIPROTKB: A0A009HA42 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 150 mM NaCl, 20 mM Tris pH 7.5 | |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Details: 60 seconds at 5 mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV Details: Blot force: 20 Blot time: 4 seconds Drain time: 0 Wait time: 0. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 5936 / Average exposure time: 3.0 sec. / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 28.96 / Target criteria: cross-correlation coefficient |
| Output model |  PDB-8t6r: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)