[English] 日本語
 Yorodumi
Yorodumi- EMDB-40660: Cryo-EM structure of human CST bound to POT1(ESDL)/TPP1 in the pr... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human CST bound to POT1(ESDL)/TPP1 in the presence of telomeric ssDNA | |||||||||
 Map data Map data | Sharpened full map of human CST bound to POT1(ESDL)/TPP1 in the presence of telomeric ssDNA | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | telomere / shelterin / cst / complex / DNA BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of single-stranded telomeric DNA binding / positive regulation of DNA strand elongation / regulation of DNA helicase activity / positive regulation of DNA helicase activity / G-rich single-stranded DNA binding / telomere assembly / CST complex / segmentation / urogenital system development / positive regulation of helicase activity ...positive regulation of single-stranded telomeric DNA binding / positive regulation of DNA strand elongation / regulation of DNA helicase activity / positive regulation of DNA helicase activity / G-rich single-stranded DNA binding / telomere assembly / CST complex / segmentation / urogenital system development / positive regulation of helicase activity / 8-hydroxy-2'-deoxyguanosine DNA binding / regulation of double-strand break repair via nonhomologous end joining / telomeric D-loop binding / protection from non-homologous end joining at telomere / telomerase inhibitor activity / DEAD/H-box RNA helicase binding / establishment of protein localization to telomere / telomeric D-loop disassembly / telomere maintenance via telomere lengthening / shelterin complex / Telomere C-strand synthesis initiation / regulation of telomere maintenance via telomerase / telomere capping / Telomere C-strand (Lagging Strand) Synthesis / : / single-stranded telomeric DNA binding / positive regulation of telomere maintenance / nuclear telomere cap complex / Processive synthesis on the C-strand of the telomere / Polymerase switching on the C-strand of the telomere / Removal of the Flap Intermediate from the C-strand / G-rich strand telomeric DNA binding / telomerase holoenzyme complex / embryonic limb morphogenesis / bone marrow development / intermediate filament cytoskeleton / protein localization to chromosome, telomeric region / DNA duplex unwinding / hematopoietic stem cell proliferation / telomeric DNA binding / : / negative regulation of telomere maintenance via telomerase / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / telomere maintenance via telomerase / Telomere Extension By Telomerase / replicative senescence / carbohydrate transmembrane transporter activity / DNA polymerase binding / Packaging Of Telomere Ends / spleen development / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / positive regulation of telomere maintenance via telomerase / regulation of G2/M transition of mitotic cell cycle / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / telomere maintenance / Meiotic synapsis / picornain 2A / protein complex oligomerization / symbiont-mediated suppression of host mRNA export from nucleus / bioluminescence / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / monoatomic ion channel activity / thymus development / positive regulation of DNA replication / T=pseudo3 icosahedral viral capsid / skeletal system development / generation of precursor metabolites and energy / intracellular protein transport / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane / multicellular organism growth / DNA Damage/Telomere Stress Induced Senescence / endocytosis involved in viral entry into host cell / fibrillar center / positive regulation of fibroblast proliferation / nucleoside-triphosphate phosphatase / single-stranded DNA binding / outer membrane-bounded periplasmic space / chromosome, telomeric region / RNA helicase activity / nuclear body / induction by virus of host autophagy / RNA-directed RNA polymerase / viral RNA genome replication / cysteine-type endopeptidase activity / intracellular membrane-bounded organelle / RNA-dependent RNA polymerase activity / virus-mediated perturbation of host defense response / DNA-templated transcription / DNA damage response / protein-containing complex binding / virion attachment to host cell / structural molecule activity / ATP hydrolysis activity / proteolysis / RNA binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
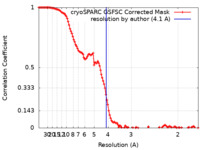
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Cai SW | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation | Journal: bioRxiv / Year: 2023 Title: POT1 recruits and regulates CST-Polα/Primase at human telomeres. Authors: Sarah W Cai / Hiroyuki Takai / Thomas Walz / Titia de Lange /  Abstract: Telomere maintenance requires extension of the G-rich telomeric repeat strand by telomerase and fill-in synthesis of the C-rich strand by Polα/Primase. Telomeric Polα/Primase is bound to Ctc1-Stn1- ...Telomere maintenance requires extension of the G-rich telomeric repeat strand by telomerase and fill-in synthesis of the C-rich strand by Polα/Primase. Telomeric Polα/Primase is bound to Ctc1-Stn1-Ten1 (CST), a single-stranded DNA-binding complex. Like mutations in telomerase, mutations affecting CST-Polα/Primase result in pathological telomere shortening and cause a telomere biology disorder, Coats plus (CP). We determined cryogenic electron microscopy structures of human CST bound to the shelterin heterodimer POT1/TPP1 that reveal how CST is recruited to telomeres by POT1. Phosphorylation of POT1 is required for CST recruitment, and the complex is formed through conserved interactions involving several residues mutated in CP. Our structural and biochemical data suggest that phosphorylated POT1 holds CST-Polα/Primase in an inactive auto-inhibited state until telomerase has extended the telomere ends. We propose that dephosphorylation of POT1 releases CST-Polα/Primase into an active state that completes telomere replication through fill-in synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40660.map.gz emd_40660.map.gz | 157 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40660-v30.xml emd-40660-v30.xml emd-40660.xml emd-40660.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40660_fsc.xml emd_40660_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_40660.png emd_40660.png | 118 KB | ||
| Masks |  emd_40660_msk_1.map emd_40660_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40660.cif.gz emd-40660.cif.gz | 8.2 KB | ||
| Others |  emd_40660_half_map_1.map.gz emd_40660_half_map_1.map.gz emd_40660_half_map_2.map.gz emd_40660_half_map_2.map.gz | 154.6 MB 154.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40660 http://ftp.pdbj.org/pub/emdb/structures/EMD-40660 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40660 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40660 | HTTPS FTP |
-Validation report
| Summary document |  emd_40660_validation.pdf.gz emd_40660_validation.pdf.gz | 880.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40660_full_validation.pdf.gz emd_40660_full_validation.pdf.gz | 880 KB | Display | |
| Data in XML |  emd_40660_validation.xml.gz emd_40660_validation.xml.gz | 20.3 KB | Display | |
| Data in CIF |  emd_40660_validation.cif.gz emd_40660_validation.cif.gz | 26.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40660 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40660 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40660 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40660 | HTTPS FTP |
-Related structure data
| Related structure data |  8sokMC  8sojC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40660.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40660.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened full map of human CST bound to POT1(ESDL)/TPP1 in the presence of telomeric ssDNA | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40660_msk_1.map emd_40660_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B of human CST bound to...
| File | emd_40660_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of human CST bound to POT1(ESDL)/TPP1 in the presence of telomeric ssDNA | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A of human CST bound to...
| File | emd_40660_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A of human CST bound to POT1(ESDL)/TPP1 in the presence of telomeric ssDNA | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ssDNA-bound human CST-POT1(ESDL)/TPP1 complex
| Entire | Name: ssDNA-bound human CST-POT1(ESDL)/TPP1 complex |
|---|---|
| Components |
|
-Supramolecule #1: ssDNA-bound human CST-POT1(ESDL)/TPP1 complex
| Supramolecule | Name: ssDNA-bound human CST-POT1(ESDL)/TPP1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 / Details: CST-POT1(ESDL)/TPP1 complex in the presence of DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 370 KDa |
-Macromolecule #1: CST complex subunit CTC1
| Macromolecule | Name: CST complex subunit CTC1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 178.300609 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGSSHHHHHH SSGTKIEEGK LVIWINGDKG YNGLAEVGKK FEKDTGIKVT VEHPDKLEEK FPQVAATGDG PDIIFWAHDR FGGYAQSGL LAEITPDKAF QDKLYPFTWD AVRYNGKLIA YPIAVEALSL IYNKDLLPNP PKTWEEIPAL DKELKAKGKS A LMFNLQEP ...String: MGSSHHHHHH SSGTKIEEGK LVIWINGDKG YNGLAEVGKK FEKDTGIKVT VEHPDKLEEK FPQVAATGDG PDIIFWAHDR FGGYAQSGL LAEITPDKAF QDKLYPFTWD AVRYNGKLIA YPIAVEALSL IYNKDLLPNP PKTWEEIPAL DKELKAKGKS A LMFNLQEP YFTWPLIAAD GGYAFKYENG KYDIKDVGVD NAGAKAGLTF LVDLIKNKHM NADTDYSIAE AAFNKGETAM TI NGPWAWS NIDTSKVNYG VTVLPTFKGQ PSKPFVGVLS AGINAASPNK ELAKEFLENY LLTDEGLEAV NKDKPLGAVA LKS YEEELA KDPRIAATME NAQKGEIMPN IPQMSAFWYA VRTAVINAAS GRQTVDEALK DAQTKLVEKY LEVLFQGPGS MAAG RAQVP SSEQAWLEDA QVFIQKTLCP AVKEPNVQLT PLVIDCVKTV WLSQGRNQGS TLPLSYSFVS VQDLKTHQRL PCCSH LSWS SSAYQAWAQE AGPNGNPLPR EQLLLLGTLT DLSADLEQEC RNGSLYVRDN TGVLSCELID LDLSWLGHLF LFPRWS YLP PARWNSSGEG HLELWDAPVP VFPLTISPGP VTPIPVLYPE SASCLLRLRN KLRGVQRNLA GSLVRLSALV KSKQKAY FI LSLGRSHPAV THVSIIVQVP AQLVWHRALR PGTAYVLTEL RVSKIRGQRQ HVWMTSQSSR LLLLKPECVQ ELELELEG P LLEADPKPLP MPSNSEDKKD PESLVRYSRL LSYSGAVTGV LNEPAGLYEL DGQLGLCLAY QQFRGLRRVM RPGVCLQLQ DVHLLQSVGG GTRRPVLAPC LRGAVLLQSF SRQKPGAHSS RQAYGASLYE QLVWERQLGL PLYLWATKAL EELACKLCPH VLRHHQFLQ HSSPGSPSLG LQLLAPTLDL LAPPGSPVRN AHNEILEEPH HCPLQKYTRL QTPSSFPTLA TLKEEGQRKA W ASFDPKAL LPLPEASYLP SCQLNRRLAW SWLCLLPSAF CPAQVLLGVL VASSHKGCLQ LRDQSGSLPC LLLAKHSQPL SD PRLIGCL VRAERFQLIV ERDVRSSFPS WKELSMPGFI QKQQARVYVQ FFLADALILP VPRPCLHSAT PSTPQTDPTG PEG PHLGQS RLFLLCHKEA LMKRNFCVPP GASPEVPKPA LSFYVLGSWL GGTQRKEGTG WGLPEPQGND DNDQKVHLIF FGSS VRWFE FLHPGQVYRL IAPGPATPML FEKDGSSCIS RRPLELAGCA SCLTVQDNWT LELESSQDIQ DVLDANKSLP ESSLT DLLS DNFTDSLVSF SAEILSRTLC EPLVASLWMK LGNTGAMRRC VKLTVALETA ECEFPPHLDV YIEDPHLPPS LGLLPG ARV HFSQLEKRVS RSHNVYCCFR SSTYVQVLSF PPETTISIPL PHIYLAELLQ GGQSPFQATA SCHIVSVFSL QLFWVCA YC TSICRQGKCT RLGSTCPTQT AISQAIIRLL VEDGTAEAVV TCRNHHVAAA LGLCPREWAS LLDFVQVPGR VVLQFAGP G AQLESSARVD EPMTMFLWTL CTSPSVLRPI VLSFELERKP SKIVPLEPPR LQRFQCGELP FLTHVNPRLR LSCLSIRES EYSSSLGILA SSC UniProtKB: Maltose/maltodextrin-binding periplasmic protein, CST complex subunit CTC1 |
-Macromolecule #2: CST complex subunit STN1
| Macromolecule | Name: CST complex subunit STN1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.172949 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MQPGSSRCEE ETPSLLWGLD PVFLAFAKLY IRDILDMKES RQVPGVFLYN GHPIKQVDVL GTVIGVRERD AFYSYGVDDS TGVINCICW KKLNTESVSA APSAARELSL TSQLKKLQET IEQKTKIEIG DTIRVRGSIR TYREEREIHA TTYYKVDDPV W NIQIARML ...String: MQPGSSRCEE ETPSLLWGLD PVFLAFAKLY IRDILDMKES RQVPGVFLYN GHPIKQVDVL GTVIGVRERD AFYSYGVDDS TGVINCICW KKLNTESVSA APSAARELSL TSQLKKLQET IEQKTKIEIG DTIRVRGSIR TYREEREIHA TTYYKVDDPV W NIQIARML ELPTIYRKVY DQPFHSSALE KEEALSNPGA LDLPSLTSLL SEKAKEFLME NRVQSFYQQE LEMVESLLSL AN QPVIHSA SSDQVNFKKD TTSKAIHSIF KNAIQLLQEK GLVFQKDDGF DNLYYVTRED KDLHRKIHRI IQQDCQKPNH MEK GCHFLH ILACARLSIR PGLSEAVLQQ VLELLEDQSD IVSTMEHYYT AF UniProtKB: CST complex subunit STN1 |
-Macromolecule #3: CST complex subunit TEN1
| Macromolecule | Name: CST complex subunit TEN1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.872013 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MMLPKPGTYY LPWEVSAGQV PDGSTLRTFG RLCLYDMIQS RVTLMAQHGS DQHQVLVCTK LVEPFHAQVG SLYIVLGELQ HQQDRGSVV KARVLTCVEG MNLPLLEQAI REQRLYKQER GGSQ UniProtKB: CST complex subunit TEN1 |
-Macromolecule #4: Protection of telomeres protein 1
| Macromolecule | Name: Protection of telomeres protein 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 72.938219 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MHHHHHHGSS LVPATNYIYT PLNQLKGGTI VNVYGVVKFF KPPYLSKGTD YCSVVTIVDQ TNVKLTCLLF SGNYEALPII YKNGDIVRF HRLKIQVYKK ETQGITSSGF ASLTFEGTLG APIIPRTSSK YFNFTTEDHK MVEALRVWAS THMSPSWTLL K LCDVQPMQ ...String: MHHHHHHGSS LVPATNYIYT PLNQLKGGTI VNVYGVVKFF KPPYLSKGTD YCSVVTIVDQ TNVKLTCLLF SGNYEALPII YKNGDIVRF HRLKIQVYKK ETQGITSSGF ASLTFEGTLG APIIPRTSSK YFNFTTEDHK MVEALRVWAS THMSPSWTLL K LCDVQPMQ YFDLTCQLLG KAEVDGASFL LKVWDGTRTP FPSWRVLIQD LVLEGDLSHI HRLQNLTIDI LVYDNHVHVA RS LKVGSFL RIYSLHTKLQ SMNSENQTML SLEFHLHGGT SYGRGIRVLP ESNSDVDQLK KDLESANLTA NQHSDVICQS EPD DSFPSS GSESDLVSLY EVERCQQLSA TILTDHQYLE RTPLCAILKQ KAPQQYRIRA KLRSYKPRRL FQSVKLHCPK CHLL QEVPH EGDLDIIFQD GATKTPDVKL QNTSLYDSKI WTTKNQKGRK VAVHFVKNNG ILPLSNECLL LIEGGTLSEI CKLSN KFNS VIPVRSGHED LELLDLSAPF LIQGTIHHYG CKQCSSLRSI QNLNSLVDKT SWIPSSVAEA LGIVPLQYVF VMTFTL DDG TGVLEAYLMD SDKFFQIPAS EVLMDDDLQK SVDMIMDMFC PPGIKIDAYP WLECFIKSYN VTNGTDNQIC YQIFDTT VA EDVI UniProtKB: Protection of telomeres protein 1 |
-Macromolecule #5: TPP1
| Macromolecule | Name: TPP1 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO / EC number: picornain 2A |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.064863 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MWSHPQFEKG GGSGGGSGGS AWSHPQFEKG SVSKGEELFT GVVPILVELD GDVNGHKFSV SGEGEGDATY GKLTLKFICT TGKLPVPWP TLVTTLTYGV QCFSRYPDHM KQHDFFKSAM PEGYVQERTI FFKDDGNYKT RAEVKFEGDT LVNRIELKGI D FKEDGNIL ...String: MWSHPQFEKG GGSGGGSGGS AWSHPQFEKG SVSKGEELFT GVVPILVELD GDVNGHKFSV SGEGEGDATY GKLTLKFICT TGKLPVPWP TLVTTLTYGV QCFSRYPDHM KQHDFFKSAM PEGYVQERTI FFKDDGNYKT RAEVKFEGDT LVNRIELKGI D FKEDGNIL GHKLEYNYNS HNVYIMADKQ KNGIKVNFKI RHNIEDGSVQ LADHYQQNTP IGDGPVLLPD NHYLSTQSAL SK DPNEKRD HMVLLEFVTA AGITLGMDEL YKENLYFQGG SMAGSGRLVL RPWIRELILG SETPSSPRAG QLLEVLQDAE AAV AGPSHA PDTSDVGATL LVSDGTHSVR CLVTREALDT SDWEEKEFGF RGTEGRLLLL QDCGVHVQVA EGGAPAEFYL QVDR FSLLP TEQPRLRVPG CNQDLDVQKK LYDCLEEHLS ESTSSNAGLS LSQLLDEMRE DQEHQGALVC LAESCLTLEG PCTAP PVTH WAASRCKATG EAVYTVPSSM LCISENDQLI LSSLGPCQRT QGPEL UniProtKB: Genome polyprotein, Adrenocortical dysplasia protein homolog |
-Macromolecule #6: DNA (5'-D(*TP*TP*AP*GP*GP*GP*TP*TP*AP*G)-3')
| Macromolecule | Name: DNA (5'-D(*TP*TP*AP*GP*GP*GP*TP*TP*AP*G)-3') / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 5.682672 KDa |
| Sequence | String: (DG)(DG)(DT)(DT)(DA)(DG)(DG)(DG)(DT)(DT) (DA)(DG)(DG)(DG)(DT)(DT)(DA)(DG) |
-Macromolecule #7: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 7 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.02 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)