+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa | |||||||||
 Map data Map data | cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (composite structure) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Electron transport chain / Supercomplex / Pseudomonas aeruginosa / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytochrome-c oxidase / : / respiratory chain complex III / oxidative phosphorylation / cytochrome-c oxidase / quinol-cytochrome-c reductase / ubiquinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / electron transport coupled proton transport / : ...cytochrome-c oxidase / : / respiratory chain complex III / oxidative phosphorylation / cytochrome-c oxidase / quinol-cytochrome-c reductase / ubiquinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / electron transport coupled proton transport / : / respiratory electron transport chain / proton transmembrane transport / 2 iron, 2 sulfur cluster binding / periplasmic space / electron transfer activity / oxidoreductase activity / iron ion binding / heme binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Di Trani JM / Rubinstein JL | |||||||||
| Funding support |  Canada, 2 items Canada, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Structure of the - respiratory supercomplex from . Authors: Justin M Di Trani / Andreea A Gheorghita / Madison Turner / Peter Brzezinski / Pia Ädelroth / Siavash Vahidi / P Lynne Howell / John L Rubinstein /   Abstract: Energy conversion by electron transport chains occurs through the sequential transfer of electrons between protein complexes and intermediate electron carriers, creating the proton motive force that ...Energy conversion by electron transport chains occurs through the sequential transfer of electrons between protein complexes and intermediate electron carriers, creating the proton motive force that enables ATP synthesis and membrane transport. These protein complexes can also form higher order assemblies known as respiratory supercomplexes (SCs). The electron transport chain of the opportunistic pathogen is closely linked with its ability to invade host tissue, tolerate harsh conditions, and resist antibiotics but is poorly characterized. Here, we determine the structure of a SC that forms between the quinol:cytochrome oxidoreductase (cytochrome ) and one of the organism's terminal oxidases, cytochrome , which is found only in some bacteria. Remarkably, the SC structure also includes two intermediate electron carriers: a diheme cytochrome and a single heme cytochrome . Together, these proteins allow electron transfer from ubiquinol in cytochrome to oxygen in cytochrome . We also present evidence that different isoforms of cytochrome can participate in formation of this SC without changing the overall SC architecture. Incorporating these different subunit isoforms into the SC would allow the bacterium to adapt to different environmental conditions. Bioinformatic analysis focusing on structural motifs in the SC suggests that cytochrome - SCs also exist in other bacterial pathogens. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40601.map.gz emd_40601.map.gz | 106.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40601-v30.xml emd-40601-v30.xml emd-40601.xml emd-40601.xml | 41.3 KB 41.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40601.png emd_40601.png | 117.8 KB | ||
| Filedesc metadata |  emd-40601.cif.gz emd-40601.cif.gz | 7.8 KB | ||
| Others |  emd_40601_additional_1.map.gz emd_40601_additional_1.map.gz emd_40601_additional_2.map.gz emd_40601_additional_2.map.gz emd_40601_additional_3.map.gz emd_40601_additional_3.map.gz emd_40601_additional_4.map.gz emd_40601_additional_4.map.gz emd_40601_additional_5.map.gz emd_40601_additional_5.map.gz emd_40601_additional_6.map.gz emd_40601_additional_6.map.gz emd_40601_additional_7.map.gz emd_40601_additional_7.map.gz emd_40601_additional_8.map.gz emd_40601_additional_8.map.gz emd_40601_half_map_1.map.gz emd_40601_half_map_1.map.gz emd_40601_half_map_2.map.gz emd_40601_half_map_2.map.gz | 70.3 MB 131.7 MB 106.9 MB 70.5 MB 1.9 MB 131.8 MB 66.4 MB 107.2 MB 129.7 MB 129.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40601 http://ftp.pdbj.org/pub/emdb/structures/EMD-40601 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40601 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40601 | HTTPS FTP |
-Validation report
| Summary document |  emd_40601_validation.pdf.gz emd_40601_validation.pdf.gz | 898.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40601_full_validation.pdf.gz emd_40601_full_validation.pdf.gz | 897.9 KB | Display | |
| Data in XML |  emd_40601_validation.xml.gz emd_40601_validation.xml.gz | 14.1 KB | Display | |
| Data in CIF |  emd_40601_validation.cif.gz emd_40601_validation.cif.gz | 16.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40601 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40601 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40601 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40601 | HTTPS FTP |
-Related structure data
| Related structure data |  8smrMC  8snhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40601.map.gz / Format: CCP4 / Size: 139.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40601.map.gz / Format: CCP4 / Size: 139.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (composite structure) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||
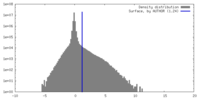
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Additional map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1 complex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1 complex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1 complex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa
+Additional map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (Locally refined...
+Additional map: cytochrome bc1 complex from Pseudomonas aeruginosa
+Half map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (Half map...
+Half map: cytochrome bc1-cbb3 supercomplex from Pseudomonas aeruginosa (Half map...
- Sample components
Sample components
+Entire : cytochrome bc1-cbb3 supercomplex
+Supramolecule #1: cytochrome bc1-cbb3 supercomplex
+Macromolecule #1: Ubiquinol-cytochrome c reductase iron-sulfur subunit
+Macromolecule #2: Cytochrome b
+Macromolecule #3: Cytochrome c1
+Macromolecule #4: Cytochrome c4
+Macromolecule #5: Cytochrome C5
+Macromolecule #6: cytochrome-c oxidase
+Macromolecule #7: Cbb3-type Cytochrome C oxidase subunit II
+Macromolecule #8: Cbb3-type cytochrome c oxidase subunit
+Macromolecule #9: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #10: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #11: UBIQUINONE-10
+Macromolecule #12: (2R)-2-(methoxymethyl)-4-{[(25R)-spirost-5-en-3beta-yl]oxy}butyl ...
+Macromolecule #13: HEME C
+Macromolecule #14: COPPER (II) ION
+Macromolecule #15: CALCIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 51.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.7 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 48594 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)