[English] 日本語
 Yorodumi
Yorodumi- EMDB-40355: Structure of rat organic anion transporter 1 (OAT1) in complex wi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of rat organic anion transporter 1 (OAT1) in complex with probenecid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | organic anion transporter / OAT / SLC22 / drug transporter / probenecid / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationOrganic anion transport by SLC22 transporters / renal tubular secretion / alpha-ketoglutarate transport / alpha-ketoglutarate transmembrane transporter activity / sodium-independent organic anion transport / : / metanephric proximal tubule development / prostaglandin transport / prostaglandin transmembrane transporter activity / solute:inorganic anion antiporter activity ...Organic anion transport by SLC22 transporters / renal tubular secretion / alpha-ketoglutarate transport / alpha-ketoglutarate transmembrane transporter activity / sodium-independent organic anion transport / : / metanephric proximal tubule development / prostaglandin transport / prostaglandin transmembrane transporter activity / solute:inorganic anion antiporter activity / organic anion transport / : / monoatomic anion transport / chloride ion binding / antiporter activity / xenobiotic transmembrane transporter activity / transmembrane transporter activity / basal plasma membrane / caveola / basolateral plasma membrane / protein-containing complex / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.86 Å | |||||||||
 Authors Authors | Dou T / Jiang J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: The substrate and inhibitor binding mechanism of polyspecific transporter OAT1 revealed by high-resolution cryo-EM. Authors: Tongyi Dou / Tengfei Lian / Shi Shu / Yi He / Jiansen Jiang /   Abstract: Organic anion transporters (OATs) of the SLC22 family have crucial roles in the transport of organic anions, including metabolites and therapeutic drugs, and in transporter-mediated drug-drug ...Organic anion transporters (OATs) of the SLC22 family have crucial roles in the transport of organic anions, including metabolites and therapeutic drugs, and in transporter-mediated drug-drug interactions. In the kidneys, OATs facilitate the elimination of metabolic waste products and xenobiotics. However, their transport activities can lead to the accumulation of certain toxic compounds within cells, causing kidney damage. Moreover, OATs are important drug targets, because their inhibition modulates the elimination or retention of substrates linked to diseases. Despite extensive research on OATs, the molecular basis of their substrate and inhibitor binding remains poorly understood. Here we report the cryo-EM structures of rat OAT1 (also known as SLC22A6) and its complexes with para-aminohippuric acid and probenecid at 2.1, 2.8 and 2.9 Å resolution, respectively. Our findings reveal a highly conserved substrate binding mechanism for SLC22 transporters, wherein four aromatic residues form a cage to accommodate the polyspecific binding of diverse compounds. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40355.map.gz emd_40355.map.gz | 14.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40355-v30.xml emd-40355-v30.xml emd-40355.xml emd-40355.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40355_fsc.xml emd_40355_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_40355.png emd_40355.png | 81.5 KB | ||
| Masks |  emd_40355_msk_1.map emd_40355_msk_1.map | 15.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40355.cif.gz emd-40355.cif.gz | 5.9 KB | ||
| Others |  emd_40355_half_map_1.map.gz emd_40355_half_map_1.map.gz emd_40355_half_map_2.map.gz emd_40355_half_map_2.map.gz | 14.5 MB 14.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40355 http://ftp.pdbj.org/pub/emdb/structures/EMD-40355 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40355 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40355 | HTTPS FTP |
-Related structure data
| Related structure data |  8sdzMC  8sduC  8sdyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40355.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40355.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40355_msk_1.map emd_40355_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_40355_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_40355_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Rat OAT1/probenecid complex solubilized in detergent LMNG
| Entire | Name: Rat OAT1/probenecid complex solubilized in detergent LMNG |
|---|---|
| Components |
|
-Supramolecule #1: Rat OAT1/probenecid complex solubilized in detergent LMNG
| Supramolecule | Name: Rat OAT1/probenecid complex solubilized in detergent LMNG type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61 KDa |
-Macromolecule #1: Solute carrier family 22 member 6
| Macromolecule | Name: Solute carrier family 22 member 6 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 60.816836 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAFNDLLKQV GGVGRFQLIQ VTMVVAPLLL MASHNTLQNF TAAIPPHHCR PPANANLSKD GGLEAWLPLD KQGQPESCLR FTSPQWGPP FYNGTEANGT RVTEPCIDGW VYDNSTFPST IVTEWNLVCS HRAFRQLAQS LYMVGVLLGA MVFGYLADRL G RRKVLILN ...String: MAFNDLLKQV GGVGRFQLIQ VTMVVAPLLL MASHNTLQNF TAAIPPHHCR PPANANLSKD GGLEAWLPLD KQGQPESCLR FTSPQWGPP FYNGTEANGT RVTEPCIDGW VYDNSTFPST IVTEWNLVCS HRAFRQLAQS LYMVGVLLGA MVFGYLADRL G RRKVLILN YLQTAVSGTC AAYAPNYTVY CVFRLLSGMS LASIAINCMT LNVEWMPIHT RAYVGTLIGY VYSLGQFLLA GI AYAVPHW RHLQLVVSVP FFIAFIYSWF FIESARWYSS SGRLDLTLRA LQRVARINGK QEEGAKLSIE VLRTSLQKEL TLS KGQASA MELLRCPTLR HLFLCLSMLW FATSFAYYGL VMDLQGFGVS MYLIQVIFGA VDLPAKFVCF LVINSMGRRP AQMA SLLLA GICILVNGII PKSHTIIRTS LAVLGKGCLA SSFNCIFLYT GELYPTVIRQ TGLGMGSTMA RVGSIVSPLV SMTAE FYPS MPLFIFGAVP VVASAVTALL PETLGQPLPD TVQDLKSRSR GKQNQQQQEQ QKQMMPLQAS TQEKNGL UniProtKB: Solute carrier family 22 member 6 |
-Macromolecule #2: 4-(dipropylsulfamoyl)benzoic acid
| Macromolecule | Name: 4-(dipropylsulfamoyl)benzoic acid / type: ligand / ID: 2 / Number of copies: 1 / Formula: RTO |
|---|---|
| Molecular weight | Theoretical: 285.359 Da |
| Chemical component information | 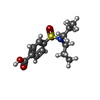 ChemComp-RTO: |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 22 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Details: TBS buffer (20 mM Tris-HCl, 150 mM NaCl, pH 8.0) with 0.01% LMNG |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 4160 / Average exposure time: 6.25 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8sdz: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)













































