[English] 日本語
 Yorodumi
Yorodumi- EMDB-40088: HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV Envelope / SOSIP / broadly neutralizing antibody / clade C / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
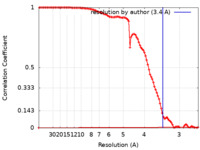
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Ozorowski G / Lee JH / Ward AB | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2023 Journal: PLoS Pathog / Year: 2023Title: Glycan heterogeneity as a cause of the persistent fraction in HIV-1 neutralization. Authors: Rajesh P Ringe / Philippe Colin / Gabriel Ozorowski / Joel D Allen / Anila Yasmeen / Gemma E Seabright / Jeong Hyun Lee / Aleksandar Antanasijevic / Kimmo Rantalainen / Thomas Ketas / John P ...Authors: Rajesh P Ringe / Philippe Colin / Gabriel Ozorowski / Joel D Allen / Anila Yasmeen / Gemma E Seabright / Jeong Hyun Lee / Aleksandar Antanasijevic / Kimmo Rantalainen / Thomas Ketas / John P Moore / Andrew B Ward / Max Crispin / P J Klasse /   Abstract: Neutralizing antibodies (NAbs) to multiple epitopes on the HIV-1-envelope glycoprotein (Env) have been isolated from infected persons. The potency of NAbs is measured more often than the size of the ...Neutralizing antibodies (NAbs) to multiple epitopes on the HIV-1-envelope glycoprotein (Env) have been isolated from infected persons. The potency of NAbs is measured more often than the size of the persistent fraction of infectivity at maximum neutralization, which may also influence preventive efficacy of active or passive immunization and the therapeutic outcome of the latter. Many NAbs neutralize HIV-1 CZA97.012, a clone of a Clade-C isolate, to ~100%. But here NAb PGT151, directed to a fusion-peptide epitope, left a persistent fraction of 15%. NAb PGT145, ligating the Env-trimer apex, left no detectable persistent fraction. The divergence in persistent fractions was further analyzed by depletion of pseudoviral populations of the most PGT151- and PGT145-reactive virions. Thereby, neutralization by the non-depleting NAb increased, whereas neutralization by the depleting NAb decreased. Furthermore, depletion by PGT151 increased sensitivity to autologous neutralization by sera from rabbits immunized with soluble native-like CZA97.012 trimer: substantial persistent fractions were reduced. NAbs in these sera target epitopes comprising residue D411 at the V4-β19 transition in a defect of the glycan shield on CZA97.012 Env. NAb binding to affinity-fractionated soluble native-like CZA97.012 trimer differed commensurately with neutralization in analyses by ELISA and surface plasmon resonance. Glycan differences between PGT151- and PGT145-purified trimer fractions were then demonstrated by mass spectrometry, providing one explanation for the differential antigenicity. These differences were interpreted in relation to a new structure at 3.4-Å resolution of the soluble CZA97.012 trimer determined by cryo-electron microscopy. The trimer adopted a closed conformation, refuting apex opening as the cause of reduced PGT145 binding to the PGT151-purified form. The evidence suggests that differences in binding and neutralization after trimer purification or pseudovirus depletion with PGT145 or PGT151 are caused by variation in glycosylation, and that some glycan variants affect antigenicity through direct effects on antibody contacts, whereas others act allosterically. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40088.map.gz emd_40088.map.gz | 86 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40088-v30.xml emd-40088-v30.xml emd-40088.xml emd-40088.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40088_fsc.xml emd_40088_fsc.xml | 10.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_40088.png emd_40088.png | 58.7 KB | ||
| Masks |  emd_40088_msk_1.map emd_40088_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40088.cif.gz emd-40088.cif.gz | 7 KB | ||
| Others |  emd_40088_half_map_1.map.gz emd_40088_half_map_1.map.gz emd_40088_half_map_2.map.gz emd_40088_half_map_2.map.gz | 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40088 http://ftp.pdbj.org/pub/emdb/structures/EMD-40088 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40088 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40088 | HTTPS FTP |
-Related structure data
| Related structure data |  8gjeMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40088.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40088.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40088_msk_1.map emd_40088_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_40088_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_40088_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
| Entire | Name: CZA97.12 SOSIP.664 in complex with 3BNC117 Fab |
|---|---|
| Components |
|
-Supramolecule #1: CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
| Supramolecule | Name: CZA97.12 SOSIP.664 in complex with 3BNC117 Fab / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Molecular weight | Theoretical: 570 KDa |
-Macromolecule #1: CZA97.12 SOSIP.664 Envelope glycoprotein gp120
| Macromolecule | Name: CZA97.12 SOSIP.664 Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 56.995121 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGARVGNMW VTVYYGVPVW TDAKTTLFCA SDTKAYDREV HNVWATHACV PTDPNPQEI VLENVTENFN MWKNDMVDQM HEDIISLWDQ SLKPCVKLTP LCVTLHCTNA TFKNNVTNDM NKEIRNCSFN T TTEIRDKK ...String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGARVGNMW VTVYYGVPVW TDAKTTLFCA SDTKAYDREV HNVWATHACV PTDPNPQEI VLENVTENFN MWKNDMVDQM HEDIISLWDQ SLKPCVKLTP LCVTLHCTNA TFKNNVTNDM NKEIRNCSFN T TTEIRDKK QQGYALFYRP DIVLLKENRN NSNNSEYILI NCNASTITQA CPKVNFDPIP IHYCAPAGYA ILKCNNKTFS GK GPCNNVS TVQCTHGIKP VVSTQLLLNG SLAEKEIIIR SENLTDNVKT IIVHLNKSVE IVCTRPNNNT RKSMRIGPGQ TFY ATGDII GDIRQAYCNI SGSKWNETLK RVKEKLQENY NNNKTIKFAP SSGGDLEITT HSFNCRGEFF YCNTTRLFNN NATE DETIT LPCRIKQIIN MWQGVGRAMY APPIAGNITC KSNITGLLLV RDGGEDNKTE EIFRPGGGNM KDNWRSELYK YKVIE LKPL GIAPTGCKRR VVERRRRRR UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: CZA97.12 SOSIP.664 Envelope glycoprotein gp41
| Macromolecule | Name: CZA97.12 SOSIP.664 Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.27967 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RQLLSSIVQQ QSNLLRAPEA QQHMLKLTVW GIKQLQTRVL AIERYLKDQQ LLGIWGCSG KLICCTNVPW NSSWSNKSQT DIWNNMTWME WDREISNYTD TIYRLLEDSQ TQQEKNEKDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #3: 3BNC117 Fab heavy chain
| Macromolecule | Name: 3BNC117 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.656484 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLLQSGAA VTKPGASVRV SCEASGYNIR DYFIHWWRQA PGQGLQWVGW INPKTGQPNN PRQFQGRVSL TRHASWDFDT YSFYMDLKA LRSDDTAVYF CARQRSDYWD FDVWGSGTQV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: QVQLLQSGAA VTKPGASVRV SCEASGYNIR DYFIHWWRQA PGQGLQWVGW INPKTGQPNN PRQFQGRVSL TRHASWDFDT YSFYMDLKA LRSDDTAVYF CARQRSDYWD FDVWGSGTQV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKVEPKSC |
-Macromolecule #4: 3BNC117 kappa light chain
| Macromolecule | Name: 3BNC117 kappa light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.022658 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPSS LSASVGDTVT ITCQANGYLN WYQQRRGKAP KLLIYDGSKL ERGVPSRFSG RRWGQEYNLT INNLQPEDIA TYFCQVYEF VVPGTRLDLK RTVAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG NSQESVTEQD S KDSTYSLS ...String: DIQMTQSPSS LSASVGDTVT ITCQANGYLN WYQQRRGKAP KLLIYDGSKL ERGVPSRFSG RRWGQEYNLT INNLQPEDIA TYFCQVYEF VVPGTRLDLK RTVAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG NSQESVTEQD S KDSTYSLS STLTLSKADY EKHKVYACEV THQGLSSPVT KSFNRGEC |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 33 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Detergent added shortly before freezing | ||||||||||||
| Grid | Model: C-flat-2/2 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 1142 / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 22500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)