+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | RuvA Holliday junction DNA complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Holliday junction / tetramer / RuvA / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationHolliday junction helicase complex / Holliday junction resolvase complex / four-way junction helicase activity / four-way junction DNA binding / DNA recombination / DNA repair / ATP binding / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   Thermus thermophilus HB8 (bacteria) / Thermus thermophilus HB8 (bacteria) /  Punavirus P1 / synthetic construct (others) Punavirus P1 / synthetic construct (others) | ||||||||||||
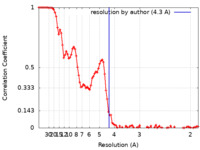
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | ||||||||||||
 Authors Authors | Rish AD / Fu T | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation | Journal: Protein Sci / Year: 2018 Title: UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Authors: Thomas D Goddard / Conrad C Huang / Elaine C Meng / Eric F Pettersen / Gregory S Couch / John H Morris / Thomas E Ferrin /  Abstract: UCSF ChimeraX is next-generation software for the visualization and analysis of molecular structures, density maps, 3D microscopy, and associated data. It addresses challenges in the size, scope, and ...UCSF ChimeraX is next-generation software for the visualization and analysis of molecular structures, density maps, 3D microscopy, and associated data. It addresses challenges in the size, scope, and disparate types of data attendant with cutting-edge experimental methods, while providing advanced options for high-quality rendering (interactive ambient occlusion, reliable molecular surface calculations, etc.) and professional approaches to software design and distribution. This article highlights some specific advances in the areas of visualization and usability, performance, and extensibility. ChimeraX is free for noncommercial use and is available from http://www.rbvi.ucsf.edu/chimerax/ for Windows, Mac, and Linux. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40036.map.gz emd_40036.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40036-v30.xml emd-40036-v30.xml emd-40036.xml emd-40036.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40036_fsc.xml emd_40036_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_40036.png emd_40036.png | 52.1 KB | ||
| Filedesc metadata |  emd-40036.cif.gz emd-40036.cif.gz | 5.5 KB | ||
| Others |  emd_40036_half_map_1.map.gz emd_40036_half_map_1.map.gz emd_40036_half_map_2.map.gz emd_40036_half_map_2.map.gz | 59.2 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40036 http://ftp.pdbj.org/pub/emdb/structures/EMD-40036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40036 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40036 | HTTPS FTP |
-Validation report
| Summary document |  emd_40036_validation.pdf.gz emd_40036_validation.pdf.gz | 736 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40036_full_validation.pdf.gz emd_40036_full_validation.pdf.gz | 735.7 KB | Display | |
| Data in XML |  emd_40036_validation.xml.gz emd_40036_validation.xml.gz | 16 KB | Display | |
| Data in CIF |  emd_40036_validation.cif.gz emd_40036_validation.cif.gz | 21 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40036 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40036 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40036 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40036 | HTTPS FTP |
-Related structure data
| Related structure data |  8gh8MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40036.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40036.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40036_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
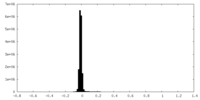
| Density Histograms |
-Half map: #1
| File | emd_40036_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
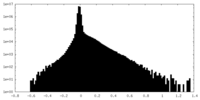
| Density Histograms |
- Sample components
Sample components
-Entire : RuvA DNA binding protein complexed with Holliday Junction DNA
| Entire | Name: RuvA DNA binding protein complexed with Holliday Junction DNA |
|---|---|
| Components |
|
-Supramolecule #1: RuvA DNA binding protein complexed with Holliday Junction DNA
| Supramolecule | Name: RuvA DNA binding protein complexed with Holliday Junction DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
-Supramolecule #2: Tetrameric RuvA binding protein
| Supramolecule | Name: Tetrameric RuvA binding protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
-Supramolecule #3: Holliday Junction DNA adapted from PDB 2HOI
| Supramolecule | Name: Holliday Junction DNA adapted from PDB 2HOI / type: complex / ID: 3 / Parent: 2 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Punavirus P1 Punavirus P1 |
-Macromolecule #1: Holliday junction branch migration complex subunit RuvA
| Macromolecule | Name: Holliday junction branch migration complex subunit RuvA type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus HB8 (bacteria) Thermus thermophilus HB8 (bacteria) |
| Molecular weight | Theoretical: 14.933622 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIRYLRGLVL KKEAGGFVLL AGGVGFFLQA PTPFLQALEE GKEVGVHTHL LLKEEGLSLY GFPDEENLAL FELLLSVSGV GPKVALALL SALPPRLLAR ALLEGDARLL TSASGVGRRL AERIALELKG KVPPHLLAGE K UniProtKB: Holliday junction branch migration complex subunit RuvA |
-Macromolecule #2: DNA (34-MER)
| Macromolecule | Name: DNA (34-MER) / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 10.399752 KDa |
| Sequence | String: (DA)(DT)(DA)(DA)(DC)(DT)(DT)(DC)(DG)(DT) (DA)(DT)(DA)(DG)(DC)(DA)(DT)(DA)(DC)(DA) (DT)(DT)(DA)(DT)(DA)(DC)(DG)(DA)(DA) (DC)(DT)(DT)(DA)(DT) |
-Macromolecule #3: DNA (34-MER)
| Macromolecule | Name: DNA (34-MER) / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 10.51081 KDa |
| Sequence | String: (DA)(DT)(DA)(DA)(DG)(DT)(DT)(DC)(DG)(DT) (DA)(DT)(DA)(DA)(DT)(DG)(DT)(DA)(DT)(DG) (DC)(DT)(DA)(DT)(DA)(DC)(DG)(DA)(DA) (DG)(DT)(DT)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)