+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TOM complex with whole Tom20 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | mitochondria / transport / membrane protein complex / PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationtRNA import into mitochondrion / TOM complex / mitochondrion targeting sequence binding / mitochondrial outer membrane translocase complex / protein insertion into mitochondrial outer membrane / mitochondria-associated endoplasmic reticulum membrane contact site / protein-transporting ATPase activity / migrasome / positive regulation of type 2 mitophagy / Mitochondrial protein import ...tRNA import into mitochondrion / TOM complex / mitochondrion targeting sequence binding / mitochondrial outer membrane translocase complex / protein insertion into mitochondrial outer membrane / mitochondria-associated endoplasmic reticulum membrane contact site / protein-transporting ATPase activity / migrasome / positive regulation of type 2 mitophagy / Mitochondrial protein import / : / : / porin activity / pore complex / protein import into mitochondrial matrix / transmembrane protein transporter activity / monoatomic ion transport / PINK1-PRKN Mediated Mitophagy / cell periphery / regulation of protein stability / intracellular protein transport / unfolded protein binding / protein transport / sperm midpiece / mitochondrial outer membrane / mitochondrial inner membrane / Ub-specific processing proteases / mitochondrion / membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.92 Å | |||||||||
 Authors Authors | Tian XY / Su JY / Sui SF | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: PNAS Nexus / Year: 2024 Journal: PNAS Nexus / Year: 2024Title: Structure of the intact Tom20 receptor in the human translocase of the outer membrane complex. Authors: Jiayue Su / Xuyang Tian / Ziyi Wang / Jiawen Yang / Shan Sun / Sen-Fang Sui /  Abstract: The translocase of the outer membrane (TOM) complex serves as the main gate for preproteins entering mitochondria and thus plays a pivotal role in sustaining mitochondrial stability. Precursor ...The translocase of the outer membrane (TOM) complex serves as the main gate for preproteins entering mitochondria and thus plays a pivotal role in sustaining mitochondrial stability. Precursor proteins, featuring amino-terminal targeting signals (presequences) or internal targeting signals, are recognized by the TOM complex receptors Tom20, Tom22, and Tom70, and then translocated into mitochondria through Tom40. By using chemical cross-linking to stabilize Tom20 in the TOM complex, this study unveils the structure of the human TOM holo complex, encompassing the intact Tom20 component, at a resolution of approximately 6 Å by cryo-electron microscopy. Our structure shows the TOM holo complex containing only one Tom20 subunit, which is located right at the center of the complex and stabilized by extensive interactions with Tom22, Tom40, and Tom6. Based on the structure, we proposed a possible translocation mode of TOM complex, by which different receptors could work simultaneously to ensure that the preproteins recognized by them are all efficiently translocated into the mitochondria. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38694.map.gz emd_38694.map.gz | 38.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38694-v30.xml emd-38694-v30.xml emd-38694.xml emd-38694.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38694.png emd_38694.png | 26.4 KB | ||
| Filedesc metadata |  emd-38694.cif.gz emd-38694.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38694 http://ftp.pdbj.org/pub/emdb/structures/EMD-38694 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38694 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38694 | HTTPS FTP |
-Related structure data
| Related structure data |  8xvaMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38694.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38694.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0825 Å | ||||||||||||||||||||||||||||||||||||
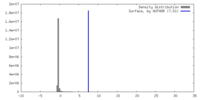
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : TOM complex with whole Tom20
| Entire | Name: TOM complex with whole Tom20 |
|---|---|
| Components |
|
-Supramolecule #1: TOM complex with whole Tom20
| Supramolecule | Name: TOM complex with whole Tom20 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 168.3 kDa/nm |
-Macromolecule #1: Mitochondrial import receptor subunit TOM6 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM6 homolog / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.007988 KDa |
| Sequence | String: MASSTVPVSA AGSANETPEI PDNVGDWLRG VYRFATDRND FRRNLILNLG LFAAGVWLAR NLSDIDLMAP QPGV UniProtKB: Mitochondrial import receptor subunit TOM6 homolog |
-Macromolecule #2: Mitochondrial import receptor subunit TOM40 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM40 homolog / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.926926 KDa |
| Sequence | String: MGNVLAASSP PAGPPPPPAP ALVGLPPPPP SPPGFTLPPL GGSLGAGTST SRSSERTPGA ATASASGAAE DGACGCLPNP GTFEECHRK CKELFPIQME GVKLTVNKGL SNHFQVNHTV ALSTIGESNY HFGVTYVGTK QLSPTEAFPV LVGDMDNSGS L NAQVIHQL ...String: MGNVLAASSP PAGPPPPPAP ALVGLPPPPP SPPGFTLPPL GGSLGAGTST SRSSERTPGA ATASASGAAE DGACGCLPNP GTFEECHRK CKELFPIQME GVKLTVNKGL SNHFQVNHTV ALSTIGESNY HFGVTYVGTK QLSPTEAFPV LVGDMDNSGS L NAQVIHQL GPGLRSKMAI QTQQSKFVNW QVDGEYRGSD FTAAVTLGNP DVLVGSGILV AHYLQSITPC LALGGELVYH RR PGEEGTV MSLAGKYTLN NWLATVTLGQ AGMHATYYHK ASDQLQVGVE FEASTRMQDT SVSFGYQLDL PKANLLFKGS VDS NWIVGA TLEKKLPPLP LTLALGAFLN HRKNKFQCGF GLTIG UniProtKB: Mitochondrial import receptor subunit TOM40 homolog |
-Macromolecule #3: Mitochondrial import receptor subunit TOM22 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM22 homolog / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.532528 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAAAVAAAGA GEPQSPDELL PKGDAEKPEE ELEEDDDEEL DETLSERLWG LTEMFPERVR SAAGATFDLS LFVAQKMYRF SRAALWIGT TSFMILVLPV VFETEKLQME QQQQLQQRQI LLGPNTGLSG GMPGALPSLP GKI UniProtKB: Mitochondrial import receptor subunit TOM22 homolog |
-Macromolecule #4: Mitochondrial import receptor subunit TOM5 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM5 homolog / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.045318 KDa |
| Sequence | String: MFRIEGLAPK LDPEEMKRKM REDVISSIRN FLIYVALLRV TPFILKKLDS I UniProtKB: Mitochondrial import receptor subunit TOM5 homolog |
-Macromolecule #5: Mitochondrial import receptor subunit TOM7 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM7 homolog / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.256473 KDa |
| Sequence | String: MVKLSKEAKQ RLQQLFKGSQ FAIRWGFIPL VIYLGFKRGA DPGMPEPTVL SLLWG UniProtKB: Mitochondrial import receptor subunit TOM7 homolog |
-Macromolecule #6: Mitochondrial import receptor subunit TOM20 homolog
| Macromolecule | Name: Mitochondrial import receptor subunit TOM20 homolog / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 16.319862 KDa |
| Sequence | String: MVGRNSAIAA GVCGALFIGY CIYFDRKRRS DPNFKNRLRE RRKKQKLAKE RAGLSKLPDL KDAEAVQKFF LEEIQLGEEL LAQGEYEKG VDHLTNAIAV CGQPQQLLQV LQQTLPPPVF QMLLTKLPTI SQRIVSAQSL AEDDVE UniProtKB: Mitochondrial import receptor subunit TOM20 homolog |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 400 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 80 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 282 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8xva: |
 Movie
Movie Controller
Controller







 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)