+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human GPR34 -Gi complex bound to M1, receptor focused | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationG protein-coupled purinergic nucleotide receptor activity / G protein-coupled receptor activity / G protein-coupled receptor signaling pathway / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
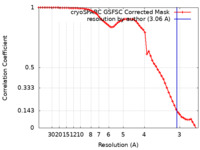
| Method | single particle reconstruction / cryo EM / Resolution: 3.06 Å | |||||||||
 Authors Authors | Kawahara R / Shihoya W / Nureki O | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis for lysophosphatidylserine recognition by GPR34. Authors: Tamaki Izume / Ryo Kawahara / Akiharu Uwamizu / Luying Chen / Shun Yaginuma / Jumpei Omi / Hiroki Kawana / Fengjue Hou / Fumiya K Sano / Tatsuki Tanaka / Kazuhiro Kobayashi / Hiroyuki H ...Authors: Tamaki Izume / Ryo Kawahara / Akiharu Uwamizu / Luying Chen / Shun Yaginuma / Jumpei Omi / Hiroki Kawana / Fengjue Hou / Fumiya K Sano / Tatsuki Tanaka / Kazuhiro Kobayashi / Hiroyuki H Okamoto / Yoshiaki Kise / Tomohiko Ohwada / Junken Aoki / Wataru Shihoya / Osamu Nureki /  Abstract: GPR34 is a recently identified G-protein coupled receptor, which has an immunomodulatory role and recognizes lysophosphatidylserine (LysoPS) as a putative ligand. Here, we report cryo-electron ...GPR34 is a recently identified G-protein coupled receptor, which has an immunomodulatory role and recognizes lysophosphatidylserine (LysoPS) as a putative ligand. Here, we report cryo-electron microscopy structures of human GPR34-G complex bound with one of two ligands bound: either the LysoPS analogue S3E-LysoPS, or M1, a derivative of S3E-LysoPS in which oleic acid is substituted with a metabolically stable aromatic fatty acid surrogate. The ligand-binding pocket is laterally open toward the membrane, allowing lateral entry of lipidic agonists into the cavity. The amine and carboxylate groups of the serine moiety are recognized by the charged residue cluster. The acyl chain of S3E-LysoPS is bent and fits into the L-shaped hydrophobic pocket in TM4-5 gap, and the aromatic fatty acid surrogate of M1 fits more appropriately. Molecular dynamics simulations further account for the LysoPS-regioselectivity of GPR34. Thus, using a series of structural and physiological experiments, we provide evidence that chemically unstable 2-acyl LysoPS is the physiological ligand for GPR34. Overall, we anticipate the present structures will pave the way for development of novel anticancer drugs that specifically target GPR34. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38219.map.gz emd_38219.map.gz | 1.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38219-v30.xml emd-38219-v30.xml emd-38219.xml emd-38219.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38219_fsc.xml emd_38219_fsc.xml | 5.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_38219.png emd_38219.png | 82.6 KB | ||
| Masks |  emd_38219_msk_1.map emd_38219_msk_1.map | 1.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-38219.cif.gz emd-38219.cif.gz | 5.7 KB | ||
| Others |  emd_38219_half_map_1.map.gz emd_38219_half_map_1.map.gz emd_38219_half_map_2.map.gz emd_38219_half_map_2.map.gz | 1.5 MB 1.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38219 http://ftp.pdbj.org/pub/emdb/structures/EMD-38219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38219 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38219 | HTTPS FTP |
-Validation report
| Summary document |  emd_38219_validation.pdf.gz emd_38219_validation.pdf.gz | 677.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_38219_full_validation.pdf.gz emd_38219_full_validation.pdf.gz | 677.4 KB | Display | |
| Data in XML |  emd_38219_validation.xml.gz emd_38219_validation.xml.gz | 9.2 KB | Display | |
| Data in CIF |  emd_38219_validation.cif.gz emd_38219_validation.cif.gz | 12.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38219 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38219 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38219 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38219 | HTTPS FTP |
-Related structure data
| Related structure data |  8xbiMC  8xbeC  8xbgC  8xbhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38219.map.gz / Format: CCP4 / Size: 1.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38219.map.gz / Format: CCP4 / Size: 1.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.291 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_38219_msk_1.map emd_38219_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
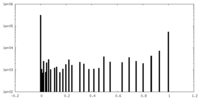
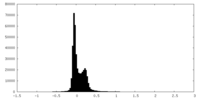
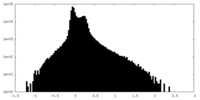
| Density Histograms |
-Half map: #2
| File | emd_38219_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_38219_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human GPR34-Gi complex bound to M1,receptor focused
| Entire | Name: Human GPR34-Gi complex bound to M1,receptor focused |
|---|---|
| Components |
|
-Supramolecule #1: Human GPR34-Gi complex bound to M1,receptor focused
| Supramolecule | Name: Human GPR34-Gi complex bound to M1,receptor focused / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Probable G-protein coupled receptor 34
| Macromolecule | Name: Probable G-protein coupled receptor 34 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.702742 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DYKDDDDAMG RSHTITMTTT SVSSWPYSSH RMRFITNHSD QPPQNFSATP NVTTCPMDEK LLSTVLTTSY SVIFIVGLVG NIIALYVFL GIHRKRNSIQ IYLLNVAIAD LLLIFCLPFR IMYHINQNKW TLGVILCKVV GTLFYMNMYI SIILLGFISL D RYIKINRS ...String: DYKDDDDAMG RSHTITMTTT SVSSWPYSSH RMRFITNHSD QPPQNFSATP NVTTCPMDEK LLSTVLTTSY SVIFIVGLVG NIIALYVFL GIHRKRNSIQ IYLLNVAIAD LLLIFCLPFR IMYHINQNKW TLGVILCKVV GTLFYMNMYI SIILLGFISL D RYIKINRS IQQRKAITTK QSIYVCCIVW MLALGGFLTM IILTLKKGGH NSTMCFHYRD KHNAKGEAIF NFILVVMFWL IF LLIILSY IKIGKNLLRI SKRRSKFPNS GKYATTARNS FIVLIIFTIC FVPYHAFRFI YISSQLNVSS CYWKEIVHKT NEI MLVLSS FNSCLDPVMY FLMSSNIRKI MCQLLFRRFQ GEPSRSESTS EFKPGYSLHD TSVAVKIQSS SKSTENLYFQ UniProtKB: Probable G-protein coupled receptor 34 |
-Macromolecule #2: (2~{S})-2-azanyl-3-[[(2~{R})-1-ethoxy-3-[3-[2-[(3-phenoxyphenyl)m...
| Macromolecule | Name: (2~{S})-2-azanyl-3-[[(2~{R})-1-ethoxy-3-[3-[2-[(3-phenoxyphenyl)methoxy]phenyl]propanoyloxy]propan-2-yl]oxy-oxidanyl-phosphoryl]oxy-propanoic acid type: ligand / ID: 2 / Number of copies: 1 / Formula: KW3 |
|---|---|
| Molecular weight | Theoretical: 617.581 Da |
| Chemical component information |  ChemComp-KW3: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 12 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z
Z Y
Y X
X