[English] 日本語
 Yorodumi
Yorodumi- EMDB-37717: Structure of monkeypox virus polymerase complex F8-A22-E4-H5 with... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of monkeypox virus polymerase complex F8-A22-E4-H5 with exgenous DNA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral DNA replication / monkeypox virus / polymerase / H5 / VIRAL PROTEIN | |||||||||
| Biological species |  Monkeypox virus / Monkeypox virus /  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Wang X / Li N / Gao N | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2023 Journal: Mol Cell / Year: 2023Title: Structural insights into the assembly and mechanism of mpox virus DNA polymerase complex F8-A22-E4-H5. Authors: Xiaohan Wang / Liangwen Ma / Ningning Li / Ning Gao /  Abstract: The DNA replication of mpox virus is performed by the viral polymerase F8 and also requires other viral factors, including processivity factor A22, uracil DNA glycosylase E4, and phosphoprotein H5. ...The DNA replication of mpox virus is performed by the viral polymerase F8 and also requires other viral factors, including processivity factor A22, uracil DNA glycosylase E4, and phosphoprotein H5. However, the molecular roles of these viral factors remain unclear. Here, we characterize the structures of F8-A22-E4 and F8-A22-E4-H5 complexes in the presence of different primer-template DNA substrates. E4 is located upstream of F8 on the template single-stranded DNA (ssDNA) and is catalytically active, highlighting a functional coupling between DNA base-excision repair and DNA synthesis. Moreover, H5, in the form of tetramer, binds to the double-stranded DNA (dsDNA) region downstream of F8 in a similar position as PCNA (proliferating cell nuclear antigen) does in eukaryotic polymerase complexes. Omission of H5 or disruption of its DNA interaction showed a reduced synthesis of full-length DNA products. These structures provide snapshots for the working cycle of the polymerase and generate insights into the mechanisms of these essential factors in viral DNA replication. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37717.map.gz emd_37717.map.gz | 10.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37717-v30.xml emd-37717-v30.xml emd-37717.xml emd-37717.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_37717.png emd_37717.png | 118 KB | ||
| Masks |  emd_37717_msk_1.map emd_37717_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-37717.cif.gz emd-37717.cif.gz | 7.1 KB | ||
| Others |  emd_37717_half_map_1.map.gz emd_37717_half_map_1.map.gz emd_37717_half_map_2.map.gz emd_37717_half_map_2.map.gz | 80.8 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37717 http://ftp.pdbj.org/pub/emdb/structures/EMD-37717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37717 | HTTPS FTP |
-Related structure data
| Related structure data |  8wpkMC  8wpeC  8wpfC  8wppC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37717.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37717.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
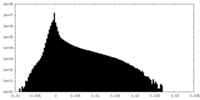
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_37717_msk_1.map emd_37717_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
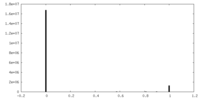
| Density Histograms |
-Half map: #2
| File | emd_37717_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
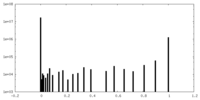
| Density Histograms |
-Half map: #1
| File | emd_37717_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
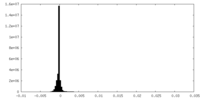
| Density Histograms |
- Sample components
Sample components
+Entire : F8-A22-E4-H5 with exogenous DNA
+Supramolecule #1: F8-A22-E4-H5 with exogenous DNA
+Supramolecule #2: F8-A22-E4-H5
+Supramolecule #3: DNA
+Macromolecule #1: DNA polymerase
+Macromolecule #2: DNA polymerase processivity factor
+Macromolecule #3: E4R Uracil-DNA glycosylase, DNA polymerase processivity factor
+Macromolecule #4: H5R late gene transcription factor
+Macromolecule #5: Primer DNA
+Macromolecule #6: Template DNA
+Macromolecule #7: 2',3'-DIDEOXY-THYMIDINE-5'-TRIPHOSPHATE
+Macromolecule #8: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 59.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.7 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1301000 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)