[English] 日本語
 Yorodumi
Yorodumi- EMDB-37656: Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate (20-nt complementary) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cas9 complex / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA-RNA complex | |||||||||
| Biological species |  Chryseobacterium (bacteria) Chryseobacterium (bacteria) | |||||||||
| Method | single particle reconstruction / Resolution: 2.98 Å | |||||||||
 Authors Authors | Zhang S / Lin S / Liu JJG | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Pro-CRISPR PcrIIC1-associated Cas9 system for enhanced bacterial immunity. Authors: Shouyue Zhang / Ao Sun / Jing-Mei Qian / Shuo Lin / Wenjing Xing / Yun Yang / Han-Zhou Zhu / Xin-Yi Zhou / Yan-Shuo Guo / Yun Liu / Yu Meng / Shu-Lin Jin / Wenhao Song / Cheng-Ping Li / ...Authors: Shouyue Zhang / Ao Sun / Jing-Mei Qian / Shuo Lin / Wenjing Xing / Yun Yang / Han-Zhou Zhu / Xin-Yi Zhou / Yan-Shuo Guo / Yun Liu / Yu Meng / Shu-Lin Jin / Wenhao Song / Cheng-Ping Li / Zhaofu Li / Shuai Jin / Jian-Hua Wang / Meng-Qiu Dong / Caixia Gao / Chunlai Chen / Yang Bai / Jun-Jie Gogo Liu /  Abstract: The CRISPR system is an adaptive immune system found in prokaryotes that defends host cells against the invasion of foreign DNA. As part of the ongoing struggle between phages and the bacterial ...The CRISPR system is an adaptive immune system found in prokaryotes that defends host cells against the invasion of foreign DNA. As part of the ongoing struggle between phages and the bacterial immune system, the CRISPR system has evolved into various types, each with distinct functionalities. Type II Cas9 is the most extensively studied of these systems and has diverse subtypes. It remains uncertain whether members of this family can evolve additional mechanisms to counter viral invasions. Here we identify 2,062 complete Cas9 loci, predict the structures of their associated proteins and reveal three structural growth trajectories for type II-C Cas9. We found that novel associated genes (NAGs) tended to be present within the loci of larger II-C Cas9s. Further investigation revealed that CbCas9 from Chryseobacterium species contains a novel β-REC2 domain, and forms a heterotetrameric complex with an NAG-encoded CRISPR-Cas-system-promoting (pro-CRISPR) protein of II-C Cas9 (PcrIIC1). The CbCas9-PcrIIC1 complex exhibits enhanced DNA binding and cleavage activity, broader compatibility for protospacer adjacent motif sequences, increased tolerance for mismatches and improved anti-phage immunity, compared with stand-alone CbCas9. Overall, our work sheds light on the diversity and 'growth evolutionary' trajectories of II-C Cas9 proteins at the structural level, and identifies many NAGs-such as PcrIIC1, which serves as a pro-CRISPR factor to enhance CRISPR-mediated immunity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37656.map.gz emd_37656.map.gz | 145.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37656-v30.xml emd-37656-v30.xml emd-37656.xml emd-37656.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_37656.png emd_37656.png | 38.9 KB | ||
| Filedesc metadata |  emd-37656.cif.gz emd-37656.cif.gz | 7.6 KB | ||
| Others |  emd_37656_half_map_1.map.gz emd_37656_half_map_1.map.gz emd_37656_half_map_2.map.gz emd_37656_half_map_2.map.gz | 151.4 MB 151.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37656 http://ftp.pdbj.org/pub/emdb/structures/EMD-37656 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37656 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37656 | HTTPS FTP |
-Related structure data
| Related structure data |  8wmmMC  8iyqC  8wmhC  8wmnC  8wr4C M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_37656.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37656.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_37656_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
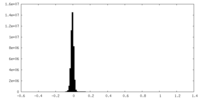
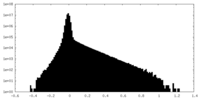
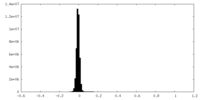
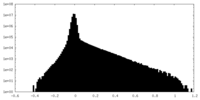
| Density Histograms |
-Half map: #1
| File | emd_37656_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate ...
| Entire | Name: Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate (20-nt complementary) |
|---|---|
| Components |
|
-Supramolecule #1: Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate ...
| Supramolecule | Name: Structure of CbCas9-PcrIIC1 complex bound to 28-bp DNA substrate (20-nt complementary) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 469.6 KDa |
-Macromolecule #1: deadCbCas9
| Macromolecule | Name: deadCbCas9 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 170.029109 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIKNILGLAL GTNSIGWALV KQDFENKQGE ILGMGSRIIP MSQDILGDFG KGNSVSQTAE RTKYRSVRRL RERFLLRRER LHRVLYILN FLPEHYASQI DFEKRLGKFK VETEPKLVWK NTDGQFSFLF QNSFNEMLED FKAAGQELKI PYDWTIYHLR K KAISQKIE ...String: MIKNILGLAL GTNSIGWALV KQDFENKQGE ILGMGSRIIP MSQDILGDFG KGNSVSQTAE RTKYRSVRRL RERFLLRRER LHRVLYILN FLPEHYASQI DFEKRLGKFK VETEPKLVWK NTDGQFSFLF QNSFNEMLED FKAAGQELKI PYDWTIYHLR K KAISQKIE KEELAWILLN FNHKRGYYQL RGEDFEEEKD KTFVRLKVDR IVDSGENVKG KILYDVYFEN GWKYDKQVVK TE DWVDRTK EFIVSESILK NGETKRTFKA VDSEKDWIAI KTKTEQEIEH SHKTVGTYIY ETLLQNPKQK IKGKLVRTIE RKF YKEELR QILEKQKEFH QELQSDDLYN DCIRELYRNN EVHQLTLRKK DFVHLFMEDI IFYQRPLRSQ KSSVSNCTLE FRKY KGENG AEHTQYLKAI PKSNPYYQEF RLWQWIFNLN LYTKDNDENV TKVFLNTTQD FENLFEFLNT RKEVDQKALL KHFKL NEKT HRWNFVEDKK YPCNETKTMI SSRLDKVENI SDDFLTRDIE QKIWHIIYSV NDKVEYEKAL KSFARKHHLD ESSFFE AFR KFPPFKSEYG SFSEKAIKKL LPLMRLGKYW NYAEIDKYSR ERIQKIITGE YDENIKDKVR EKSVHLTIEN DFQGLQL WL AQYIVYGRHS EASMIGKWNS ANDLEVFLKD FKQHSLRNPI VEQVITETLR VVKDIWLKYG NGTKDFFNEI HIELGREM K LPADDRKKLT NQITENENTN LRIKALLAEM MNDHSVENVR PFSPMQQEIL KIYEDGVLKS DIEIEDDILK ISKTAQPSS SDLKRYKLWL EQKYKSPYTG QIIPLNKLFT PEYEIEAIIP QSRYFDDSFS NKIICESAVN KLKDNYIGLG FIKQFGGTII ELGFGKSVK VFDTEEYEDF VKKHYANNRG KRNKLLMEDI PEKMIERQLN DTRYISKYIS GILSNIVRVE DGSDEGVNSK N IVPGNGKI TTQLKQDWGL NDVWNDLILP RFERMNQLTN SKDFTAWNEN HQKFLPTVPI EFSKGFSKKR IDHRHHALDA LV IACATTD HVNLLNNQSA KSDTKRYDLK KKLMKFEKVV YHHTQTGEKI EREIPKQFLK PWEKFTVDAK HNLESIIVSF KQN LRVINK ATNYYEKYVE KDGTKNKERV EQAGTNWAIR KPMHKDTVSG KVDLPWVKVP KGKILTATRK SLDSSFDLKS IGSI TDTGI QKILKNYLAF KDGNPELAFS PEGIDDLNKN IEKYNDGKPH QPINKVRVFE LGSKFQVGQT GNKKGKYVEA AKGTN LFFA VYEDEKGKRS YETIPLNEVI ERQKQGLTSV PLENEKGSRL LFDLSPNDLV YVPEIDENID SNFVFSNLNK EKISRI YKV EKTSGTECYF VRQDIAYLIK QYDAKTKIGE LESQNKLQVT MTDDRIRITD TCVKINCDRL GNINFITKEK IKQIFNE FR |
-Macromolecule #4: PcrIIC1
| Macromolecule | Name: PcrIIC1 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 15.991991 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSLDKIAIDT NILLYAYDNR DLDKQDRAVE ILLKKPFVTQ LVVFEFIKVL ERRFKMDKKE ITKLTIKLLK EVIIPLSLHR DIYNYSQFL LQRYNFGLSD ILVLSDSILN NCTILLSEDM CNGMIVDKKL KIVNPFL |
-Macromolecule #2: TS
| Macromolecule | Name: TS / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 8.450413 KDa |
| Sequence | String: (DC)(DG)(DT)(DT)(DT)(DT)(DG)(DT)(DC)(DT) (DC)(DG)(DG)(DC)(DT)(DC)(DC)(DC)(DC)(DG) (DA)(DC)(DA)(DT)(DT)(DC)(DT)(DC) |
-Macromolecule #5: NTS
| Macromolecule | Name: NTS / type: dna / ID: 5 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 8.76267 KDa |
| Sequence | String: (DG)(DA)(DG)(DA)(DA)(DT)(DG)(DT)(DC)(DG) (DG)(DG)(DG)(DA)(DG)(DC)(DC)(DG)(DA)(DG) (DA)(DC)(DA)(DA)(DA)(DA)(DC)(DG) |
-Macromolecule #3: sgRNA
| Macromolecule | Name: sgRNA / type: rna / ID: 3 / Number of copies: 2 |
|---|---|
| Source (natural) | Organism:  Chryseobacterium (bacteria) Chryseobacterium (bacteria) |
| Molecular weight | Theoretical: 40.936227 KDa |
| Sequence | String: GAGAAUGUCG GGGAGCCGAG GUUGUGAAUU GCUUUCAAAA AUUAUUGAGA AAUAAUUUUG AAAAGCAAUU CACAAUAAGG AUUAUUCCG UUGUGAAAAC AUUCAAGGCG GGGCAACUCG CCUUUUUU |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Component - Concentration: 1.0 uM / Component - Formula: NaCl / Component - Name: sodium chloride Details: 150mM NaCl, 20mM Hepes (pH=7.5), 5mM MgCl2, 1mM Tcep, 0.1% Glycerol |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)