+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a protein-RNA complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA / ribonuclease / RNA BINDING PROTEIN/RNA / RNA BINDING PROTEIN-RNA complex | |||||||||
| Biological species | unidentified (others) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.09 Å | |||||||||
 Authors Authors | Li Z | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2024 Journal: Nucleic Acids Res / Year: 2024Title: Molecular mechanism for target RNA recognition and cleavage of Cas13h. Authors: Fugen Chen / Chendi Zhang / Jialin Xue / Feng Wang / Zhuang Li /  Abstract: RNA-targeting type VI CRISPR-Cas effectors are widely used in RNA applications. Cas13h is a recently identified subtype of Cas13 ribonuclease, with strong RNA cleavage activity and robust in vivo RNA ...RNA-targeting type VI CRISPR-Cas effectors are widely used in RNA applications. Cas13h is a recently identified subtype of Cas13 ribonuclease, with strong RNA cleavage activity and robust in vivo RNA knockdown efficiency. However, little is known regarding its biochemical properties and working mechanisms. Biochemical characterization of Cas13h1 indicated that it lacks in vitro pre-crRNA processing activity and adopts a central seed. The cleavage activity of Cas13h1 is enhanced by a R(G/A) 5'-PFS, and inhibited by tag:anti-tag RNA pairing. We determined the structures of Cas13h1-crRNA binary complex at 3.1 Å and Cas13h1-crRNA-target RNA ternary complex at 3.0 Å. The ternary complex adopts an elongated architecture, and encodes a nucleotide-binding pocket within Helical-2 domain to recognize the guanosine at the 5'-end of the target RNA. Base pairing between crRNA guide and target RNA disrupts Cas13h1-guide interactions, leading to dramatic movement of HEPN domains. Upon target RNA engagement, Cas13h1 adopts a complicated activation mechanism, including separation of HEPN catalytic residues and destabilization of the active site loop and NTD domain, to get activated. Collectively, these insights expand our understanding into Cas13 effectors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37439.map.gz emd_37439.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37439-v30.xml emd-37439-v30.xml emd-37439.xml emd-37439.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
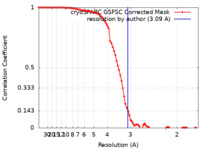
| FSC (resolution estimation) |  emd_37439_fsc.xml emd_37439_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_37439.png emd_37439.png | 88.7 KB | ||
| Filedesc metadata |  emd-37439.cif.gz emd-37439.cif.gz | 6.1 KB | ||
| Others |  emd_37439_half_map_1.map.gz emd_37439_half_map_1.map.gz emd_37439_half_map_2.map.gz emd_37439_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37439 http://ftp.pdbj.org/pub/emdb/structures/EMD-37439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37439 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37439 | HTTPS FTP |
-Validation report
| Summary document |  emd_37439_validation.pdf.gz emd_37439_validation.pdf.gz | 971.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_37439_full_validation.pdf.gz emd_37439_full_validation.pdf.gz | 971.5 KB | Display | |
| Data in XML |  emd_37439_validation.xml.gz emd_37439_validation.xml.gz | 16.4 KB | Display | |
| Data in CIF |  emd_37439_validation.cif.gz emd_37439_validation.cif.gz | 21.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37439 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37439 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37439 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37439 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_37439.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37439.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_37439_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
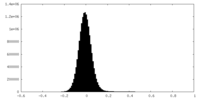
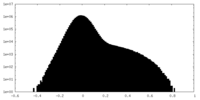
| Density Histograms |
-Half map: #1
| File | emd_37439_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
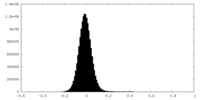
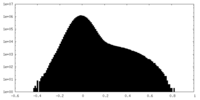
| Density Histograms |
- Sample components
Sample components
-Entire : protein-RNA complex
| Entire | Name: protein-RNA complex |
|---|---|
| Components |
|
-Supramolecule #1: protein-RNA complex
| Supramolecule | Name: protein-RNA complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism: unidentified (others) |
-Macromolecule #1: Uncharacterized protein
| Macromolecule | Name: Uncharacterized protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 130.957328 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDTPSKESSG KIIEKLDELK KATQESIEQR KKFDYRIGKG LADKLGNEKD SYFEKYLRKT VGENIPPKIL VKPKGGWKTE TKGKGKDWG DKITFAAMPQ HHYNTLLAMA FTKAHKVYSD IRGKVEIDAA QYETKAKLIE VEYGKDETRG LSGLEYLMMS K HLFSGKNS ...String: MDTPSKESSG KIIEKLDELK KATQESIEQR KKFDYRIGKG LADKLGNEKD SYFEKYLRKT VGENIPPKIL VKPKGGWKTE TKGKGKDWG DKITFAAMPQ HHYNTLLAMA FTKAHKVYSD IRGKVEIDAA QYETKAKLIE VEYGKDETRG LSGLEYLMMS K HLFSGKNS NIKLAVKKGE TEILKEYALN EKLIYTVLDE LRNFHSHIFH EPGPVSFKNL YGDEYKPEKK LTEEEWAIAR DW FVNRFND AKEHKLKTLA KVLEREGTTE EKEDAEKVIK TISGYSFEYN NCISREALLF IACMFLRKSD AAYFTKKWTG MKK AEGVFK STQSFFTDNA LKESKSILTL NADLYKYRQI LGVLSTMPAM KTDSLKPFYD FIKINNDSYS EKAEKARSKE EKEK IQAFI IPQRKSSNYT YWFMKYLNDN KLLDGFRIAY YKTPEDRFMY LIHNGLISQD DLENIEDFKT PDEKLKYLRE KGFNL KLKM KQAVGDEKKS LTEIYKETQR NFVFKVPTIE NDNFCVKKLN VFFQTDIEFN GQKISVQLSV SPDFLMKWVF VLLITG EDS IKNAITEKKE KIKDILKKYA EEYYNRCITS NFNEPLMGLE ASKVFPSSLT STVEIDEKID KDKILMRISE KYNELTK FD EENKSRKAPW RFASKRKIDI ILDYVHLVYS DRAFDEKKSV DAMRHEALND MEYMDTFEYL RYYGRYRETE EFKKIFFE D KKLYFSPILK AMKQLDSLEG VFNFAITGFL NYLKGIQSKV TDENTNKYGK VFKVTGKSLT SKIGHHSEMF SVNHCVPQE LIKLNDIKGY MKWKHETKDK LWISDFAFIR NVLESRGGFS NTDYLMKEVM PLITFEKNEK GSIKGNTQMF VALSRNKTNE LMLWEIGKY YWKEATGSEF SRLFKGLEKN TGNKITKAYR FTNPYYTIYQ EDLDIKIQRK DKKGKVIVNS PVYTIKIKPK K FDDEYQYY EQEHIVDYIE NYEPKKGIDG HWHFEELNKK IKDELARYLD DIYLLMTVEK HIVQKDFDKY ASLLKEKDSK LP PYYIGFK RNDIALNKVI FDALKHKGSF SADDLNIYRI NVLHQILQPK REKYIIIRKS LIEYCAENKL LKTM |
-Macromolecule #2: RNA (66-MER)
| Macromolecule | Name: RNA (66-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 21.257666 KDa |
| Sequence | String: UGCUUCACGU AGGCCUUGGA GCCGUACAUG GUUGUAACAA GCCUAAGUUU GAAAGGUAAA AACAAC |
-Macromolecule #3: RNA (31-MER)
| Macromolecule | Name: RNA (31-MER) / type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 9.973005 KDa |
| Sequence | String: GCAUGUACGG CUCCAAGGCC UACGUGAAGC A |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)