+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | LPS-bound P2Y10 in complex with G13 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR-G-protein complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationD5 dopamine receptor binding / P2Y receptors / regulation of fibroblast migration / Rho-activating G protein-coupled receptor signaling pathway / negative regulation of adenylate cyclase-activating adrenergic receptor signaling pathway / negative regulation of calcium ion-dependent exocytosis / G protein-coupled adenosine receptor signaling pathway / negative regulation of adenylate cyclase activity / positive regulation of neural precursor cell proliferation / negative regulation of synaptic transmission ...D5 dopamine receptor binding / P2Y receptors / regulation of fibroblast migration / Rho-activating G protein-coupled receptor signaling pathway / negative regulation of adenylate cyclase-activating adrenergic receptor signaling pathway / negative regulation of calcium ion-dependent exocytosis / G protein-coupled adenosine receptor signaling pathway / negative regulation of adenylate cyclase activity / positive regulation of neural precursor cell proliferation / negative regulation of synaptic transmission / regulation of small GTPase mediated signal transduction / positive regulation of urine volume / NRAGE signals death through JNK / branching involved in blood vessel morphogenesis / gamma-aminobutyric acid signaling pathway / regulation of calcium ion transport / CDC42 GTPase cycle / negative regulation of apoptotic signaling pathway / regulation of postsynapse assembly / neuronal dense core vesicle / Rho protein signal transduction / positive regulation of vascular associated smooth muscle cell proliferation / positive regulation of superoxide anion generation / Adenylate cyclase inhibitory pathway / RAC1 GTPase cycle / response to nutrient / guanyl-nucleotide exchange factor activity / hippocampal mossy fiber to CA3 synapse / Regulation of insulin secretion / brush border membrane / platelet activation / G protein-coupled receptor binding / G protein-coupled receptor activity / regulation of blood pressure / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / melanosome / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / regulation of cell shape / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G-protein beta-subunit binding / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / cell body / G protein activity / fibroblast proliferation / Ca2+ pathway / midbody / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / in utero embryonic development / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / cell differentiation / positive regulation of ERK1 and ERK2 cascade / cell population proliferation / Extra-nuclear estrogen signaling / postsynapse / ciliary basal body / positive regulation of cell migration Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
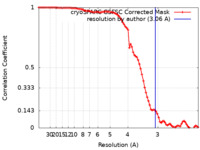
| Method | single particle reconstruction / cryo EM / Resolution: 3.06 Å | |||||||||
 Authors Authors | He Y / Yin Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Chem Biol / Year: 2024 Journal: Cell Chem Biol / Year: 2024Title: Insights into lysophosphatidylserine recognition and Gα-coupling specificity of P2Y10. Authors: Han Yin / Nozomi Kamakura / Yu Qian / Manae Tatsumi / Tatsuya Ikuta / Jiale Liang / Zhenmei Xu / Ruixue Xia / Anqi Zhang / Changyou Guo / Asuka Inoue / Yuanzheng He /   Abstract: The lysophosphatidylserine (LysoPS) receptor P2Y10, also known as LPS, plays crucial roles in the regulation of immune responses and holds promise for the treatment of autoimmune diseases. Here, we ...The lysophosphatidylserine (LysoPS) receptor P2Y10, also known as LPS, plays crucial roles in the regulation of immune responses and holds promise for the treatment of autoimmune diseases. Here, we report the cryoelectron microscopy (cryo-EM) structure of LysoPS-bound P2Y10 in complex with an engineered G heterotrimeric protein. The structure and a mutagenesis study highlight the predominant role of a comprehensive polar network in facilitating the binding and activation of the receptor by LysoPS. This interaction pattern is preserved in GPR174, but not in GPR34. Moreover, our structural study unveils the essential interactions that underlie the Gα engagement of P2Y10 and identifies key determinants for Gα-vs.-Gα-coupling selectivity, whose mutations selectively disrupt Gα engagement while preserving the intact coupling of Gα. The combined structural and functional studies provide insights into the molecular mechanisms of LysoPS recognition and Gα coupling specificity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37220.map.gz emd_37220.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37220-v30.xml emd-37220-v30.xml emd-37220.xml emd-37220.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37220_fsc.xml emd_37220_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_37220.png emd_37220.png | 84.8 KB | ||
| Filedesc metadata |  emd-37220.cif.gz emd-37220.cif.gz | 6.4 KB | ||
| Others |  emd_37220_half_map_1.map.gz emd_37220_half_map_1.map.gz emd_37220_half_map_2.map.gz emd_37220_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37220 http://ftp.pdbj.org/pub/emdb/structures/EMD-37220 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37220 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37220 | HTTPS FTP |
-Related structure data
| Related structure data |  8kggMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37220.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37220.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_37220_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_37220_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GPCR/G-protein complex
| Entire | Name: GPCR/G-protein complex |
|---|---|
| Components |
|
-Supramolecule #1: GPCR/G-protein complex
| Supramolecule | Name: GPCR/G-protein complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-2,Guanine n...
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-2,Guanine nucleotide-binding protein subunit alpha-13 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.5744 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTVSAEDK AAAERSKEID KCLSREKTYV KRLVKILLLG ADNSGKSTFL KQMRIIHGGS GGSGGTKGIH EYDFEIKNVP FKMVDVGGQ RSERKRWFEC FDSVTSILFL VDSSDFNRLT ESLNDFETIV NNRVFSNVSI ILFLNKTDLL EEKVQIVSIK D YFLEFEGD ...String: MGSTVSAEDK AAAERSKEID KCLSREKTYV KRLVKILLLG ADNSGKSTFL KQMRIIHGGS GGSGGTKGIH EYDFEIKNVP FKMVDVGGQ RSERKRWFEC FDSVTSILFL VDSSDFNRLT ESLNDFETIV NNRVFSNVSI ILFLNKTDLL EEKVQIVSIK D YFLEFEGD PHCLRDVQKF LVECFRNKRR DQQQKPLYHH FTTAINTENA RLIFRDVKDT ILHDNLKQLM LQ UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-2, Guanine nucleotide-binding protein subunit alpha-13 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.915496 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.277299 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS AGGGGSGGGG SGGGGSADIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS AGGGGSGGGG SGGGGSADIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: Putative P2Y purinoceptor 10
| Macromolecule | Name: Putative P2Y purinoceptor 10 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.812109 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MANLDKYTET FKMGSNSTST AEIYCNVTNV KFQYSLYATT YILIFIPGLL ANSAALWVLC RFISKKNKAI IFMINLSVAD LAHVLSLPL RIYYYISHHW PFQRALCLLC FYLKYLNMYA SICFLTCISL QRCFFLLKPF RARDWKRRYD VGISAAIWIV V GTACLPFP ...String: MANLDKYTET FKMGSNSTST AEIYCNVTNV KFQYSLYATT YILIFIPGLL ANSAALWVLC RFISKKNKAI IFMINLSVAD LAHVLSLPL RIYYYISHHW PFQRALCLLC FYLKYLNMYA SICFLTCISL QRCFFLLKPF RARDWKRRYD VGISAAIWIV V GTACLPFP ILRSTDLNNN KSCFADLGYK QMNAVALVGM ITVAELAGFV IPVIIIAWCT WKTTISLRQP PMAFQGISER QK ALRMVFM CAAVFFICFT PYHINFIFYT MVKETIISSC PVVRIALYFH PFCLCLASLC CLLDPILYYF MASEFRDQLS RHG SSVTRS RLMSKESGSS MIG UniProtKB: Putative P2Y purinoceptor 10 |
-Macromolecule #6: (2~{S})-2-$l^{4}-azanyl-3-[[(2~{R})-3-octadecanoyloxy-2-oxidanyl-...
| Macromolecule | Name: (2~{S})-2-$l^{4}-azanyl-3-[[(2~{R})-3-octadecanoyloxy-2-oxidanyl-propoxy]-oxidanyl-oxidanylidene-$l^{6}-phosphanyl]oxy-propanoic acid type: ligand / ID: 6 / Number of copies: 1 / Formula: WJS |
|---|---|
| Molecular weight | Theoretical: 497.391 Da |
| Chemical component information |  ChemComp-WJS: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



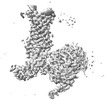































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)