+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full agonist-bound mu-type opioid receptor-G protein complex | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | SIGNALING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationOpioid Signalling / beta-endorphin receptor activity / morphine receptor activity / negative regulation of Wnt protein secretion / regulation of cellular response to stress / G protein-coupled opioid receptor signaling pathway / behavioral response to ethanol / negative regulation of nitric oxide biosynthetic process / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / sensory perception ...Opioid Signalling / beta-endorphin receptor activity / morphine receptor activity / negative regulation of Wnt protein secretion / regulation of cellular response to stress / G protein-coupled opioid receptor signaling pathway / behavioral response to ethanol / negative regulation of nitric oxide biosynthetic process / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / sensory perception / regulation of NMDA receptor activity / neuropeptide binding / GTP metabolic process / positive regulation of neurogenesis / negative regulation of cytosolic calcium ion concentration / positive regulation of macroautophagy / G-protein alpha-subunit binding / voltage-gated calcium channel activity / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / MECP2 regulates neuronal receptors and channels / sensory perception of pain / Peptide ligand-binding receptors / neuropeptide signaling pathway / electron transport chain / G protein-coupled receptor binding / G protein-coupled receptor activity / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / ADP signalling through P2Y purinoceptor 12 / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / GDP binding / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / cellular response to catecholamine stimulus / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Thrombin signalling through proteinase activated receptors (PARs) / fibroblast proliferation / Ca2+ pathway / Interleukin-4 and Interleukin-13 signaling / midbody / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / perikaryon / periplasmic space / electron transfer activity / positive regulation of ERK1 and ERK2 cascade / Extra-nuclear estrogen signaling / endosome / neuron projection / ciliary basal body / iron ion binding / G protein-coupled receptor signaling pathway / negative regulation of cell population proliferation / Golgi membrane / axon / cell division / GTPase activity / heme binding / synapse / dendrite / endoplasmic reticulum membrane / GTP binding / nucleolus / endoplasmic reticulum / Golgi apparatus / signal transduction / extracellular exosome / nucleoplasm / membrane / metal ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) / Homo sapiens (human) / synthetic construct (others) /  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.98 Å | |||||||||||||||
 Authors Authors | Uchikubo-Kamo T / Shirouzu M / Hisano T / Imai S / Kaneko S / Shimada I | |||||||||||||||
| Funding support |  Japan, 4 items Japan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural and dynamic insights into the activation of the μ-opioid receptor by an allosteric modulator. Authors: Shun Kaneko / Shunsuke Imai / Tomomi Uchikubo-Kamo / Tamao Hisano / Nobuaki Asao / Mikako Shirouzu / Ichio Shimada /  Abstract: G-protein-coupled receptors (GPCRs) play pivotal roles in various physiological processes. These receptors are activated to different extents by diverse orthosteric ligands and allosteric modulators. ...G-protein-coupled receptors (GPCRs) play pivotal roles in various physiological processes. These receptors are activated to different extents by diverse orthosteric ligands and allosteric modulators. However, the mechanisms underlying these variations in signaling activity by allosteric modulators remain largely elusive. Here, we determine the three-dimensional structure of the μ-opioid receptor (MOR), a class A GPCR, in complex with the G protein and an allosteric modulator, BMS-986122, using cryogenic electron microscopy. Our results reveal that BMS-986122 binding induces changes in the map densities corresponding to R167 and Y254, key residues in the structural motifs conserved among class A GPCRs. Nuclear magnetic resonance analyses of MOR in the absence of the G protein reveal that BMS-986122 binding enhances the formation of the interaction between R167 and Y254, thus stabilizing the fully-activated conformation, where the intracellular half of TM6 is outward-shifted to allow for interaction with the G protein. These findings illuminate that allosteric modulators like BMS-986122 can potentiate receptor activation through alterations in the conformational dynamics in the core region of GPCRs. Together, our results demonstrate the regulatory mechanisms of GPCRs, providing insights into the rational development of therapeutics targeting GPCRs. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36989.map.gz emd_36989.map.gz | 78.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36989-v30.xml emd-36989-v30.xml emd-36989.xml emd-36989.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36989_fsc.xml emd_36989_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_36989.png emd_36989.png | 76.3 KB | ||
| Filedesc metadata |  emd-36989.cif.gz emd-36989.cif.gz | 7.2 KB | ||
| Others |  emd_36989_half_map_1.map.gz emd_36989_half_map_1.map.gz emd_36989_half_map_2.map.gz emd_36989_half_map_2.map.gz | 65.2 MB 65.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36989 http://ftp.pdbj.org/pub/emdb/structures/EMD-36989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36989 | HTTPS FTP |
-Related structure data
| Related structure data |  8k9kMC  8k9lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36989.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36989.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36989_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
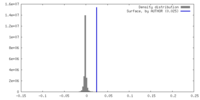
| Density Histograms |
-Half map: #1
| File | emd_36989_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Full agonist-bound mu-type opioid receptor-G protein complex
| Entire | Name: Full agonist-bound mu-type opioid receptor-G protein complex |
|---|---|
| Components |
|
-Supramolecule #1: Full agonist-bound mu-type opioid receptor-G protein complex
| Supramolecule | Name: Full agonist-bound mu-type opioid receptor-G protein complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Soluble cytochrome b562,Mu-type opioid receptor
| Macromolecule | Name: Soluble cytochrome b562,Mu-type opioid receptor / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.321855 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CLVFADYKDD DDALEVLFQG PMADLEDNWE TLNDNLKVIE KADNAAQVKD ALTKMRAAAL DAQKATPPKL EDKSPDSPEM KDFRHGFDI LVGQIDDALK LANEGKVKEA QAAAEQLKTT RNAYIQKYLG GSPGARSASS PSMITAITIM ALYSIVCVVG L FGNFLVMY ...String: CLVFADYKDD DDALEVLFQG PMADLEDNWE TLNDNLKVIE KADNAAQVKD ALTKMRAAAL DAQKATPPKL EDKSPDSPEM KDFRHGFDI LVGQIDDALK LANEGKVKEA QAAAEQLKTT RNAYIQKYLG GSPGARSASS PSMITAITIM ALYSIVCVVG L FGNFLVMY VIVRYTKMKT ATNIYIFNLA LADALATSTL PFQSVNYLMG TWPFGTILCK IVISIDYYNM FTSIWTLCTM SV DRYIAVC HPVKALDFRT PRNAKIINVC NWILSSAIGL PVMFMATTKY RQGSIDCTLT FSHPTWYWEN LLKICVFIFA FIM PVLIIT VCYGLMILRL KSVRMLSGSK EKDRNLRRIT RMVLVVVAVF IVCWTPIHIY VIIKALVTIP ETTFQTVSWH FCIA LGYTN SCLNPVLYAF LDENFKRCFR EFCIPTSSNI EQLEVLFQGP HHHHHHHH UniProtKB: Soluble cytochrome b562, Mu-type opioid receptor |
-Macromolecule #2: DAMGO
| Macromolecule | Name: DAMGO / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 513.587 Da |
| Sequence | String: Y(DAL)G(MEA)(ETA) |
-Macromolecule #3: G protein subunit alpha i3
| Macromolecule | Name: G protein subunit alpha i3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.585137 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AKEVKLLLLG AGESGKSTIV KQMKIIHEDG YSEDECKQYK VVVYSNTIQS IIAIIRAMG RLKIDFGEAA RADDARQLFV LAGSAEEGVM TPELAGVIKR LWRDGGVQAC FSRSREYQLN DSASYYLNDL D RISQSNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AKEVKLLLLG AGESGKSTIV KQMKIIHEDG YSEDECKQYK VVVYSNTIQS IIAIIRAMG RLKIDFGEAA RADDARQLFV LAGSAEEGVM TPELAGVIKR LWRDGGVQAC FSRSREYQLN DSASYYLNDL D RISQSNYI PTQQDVLRTR VKTTGIVETH FTFKDLYFKM FDVGGQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTETSIILF LNKKDLFEEK IKRSPLTICY PEYTGSNTYE EAAAYIQCQF EDLNRRKDTK EIY THFTCA TDTKNVQFVF DAVTDVIIKN NLKECGLY UniProtKB: G protein subunit alpha i3 |
-Macromolecule #4: Guanine nucleotide binding protein, beta polypeptide 1
| Macromolecule | Name: Guanine nucleotide binding protein, beta polypeptide 1 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.522098 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHHHS ELDQLRQEAE QLKNQIRDAR KACADATLSQ ITNNIDPVGR IQMRTRRTLR GHLAKIYAMH WGTDSRLLVS ASQDGKLII WDSYTTNKVH AIPLRSSWVM TCAYAPSGNY VACGGLDNIC SIYNLKTREG NVRVSRELAG HTGYLSCCRF L DDNQIVTS ...String: MHHHHHHHHS ELDQLRQEAE QLKNQIRDAR KACADATLSQ ITNNIDPVGR IQMRTRRTLR GHLAKIYAMH WGTDSRLLVS ASQDGKLII WDSYTTNKVH AIPLRSSWVM TCAYAPSGNY VACGGLDNIC SIYNLKTREG NVRVSRELAG HTGYLSCCRF L DDNQIVTS SGDTTCALWD IETGQQTTTF TGHTGDVMSL SLAPDTRLFV SGACDASAKL WDVREGMCRQ TFTGHESDIN AI CFFPNGN AFATGSDDAT CRLFDLRADQ ELMTYSHDNI ICGITSVSFS KSGRLLLAGY DDFNCNVWDA LKADRAGVLA GHD NRVSCL GVTDDGMAVA TGSWDSFLKI WN UniProtKB: Guanine nucleotide binding protein, beta polypeptide 1 |
-Macromolecule #5: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #6: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 27.409588 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAALEVLFQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 15 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average exposure time: 2.3 sec. / Average electron dose: 50.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8k9k: |
 Movie
Movie Controller
Controller































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)