+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mouse Fc epsilon RI in complex with mIgE Fc | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | IgE / high-affinity IgE receptor / allergy / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationPlatelet Adhesion to exposed collagen / IgE receptor activity / Fc-epsilon receptor I complex / serotonin secretion / Fc epsilon receptor (FCERI) signaling / Dectin-2 family / negative regulation of mast cell apoptotic process / Fc receptor mediated stimulatory signaling pathway / Role of LAT2/NTAL/LAB on calcium mobilization / T cell differentiation involved in immune response ...Platelet Adhesion to exposed collagen / IgE receptor activity / Fc-epsilon receptor I complex / serotonin secretion / Fc epsilon receptor (FCERI) signaling / Dectin-2 family / negative regulation of mast cell apoptotic process / Fc receptor mediated stimulatory signaling pathway / Role of LAT2/NTAL/LAB on calcium mobilization / T cell differentiation involved in immune response / high-affinity IgE receptor activity / GPVI-mediated activation cascade / FCERI mediated MAPK activation / mast cell apoptotic process / mast cell activation / Fc-gamma receptor III complex / FCERI mediated Ca+2 mobilization / positive regulation of interleukin-3 production / FCERI mediated NF-kB activation / serotonin secretion by platelet / positive regulation of mast cell cytokine production / Cell surface interactions at the vascular wall / neutrophil activation involved in immune response / positive regulation of mast cell degranulation / regulation of platelet activation / Fc-gamma receptor signaling pathway / positive regulation of type III hypersensitivity / IgE binding / positive regulation of type IIa hypersensitivity / regulation of release of sequestered calcium ion into cytosol / positive regulation of protein localization to cell surface / leukotriene biosynthetic process / : / positive regulation of type I hypersensitivity / interleukin-3-mediated signaling pathway / positive regulation of granulocyte macrophage colony-stimulating factor production / IgG binding / mast cell degranulation / phagocytosis, engulfment / positive regulation of interleukin-4 production / antigen processing and presentation of exogenous peptide antigen via MHC class I / Fc-epsilon receptor signaling pathway / positive regulation of interleukin-10 production / cellular response to low-density lipoprotein particle stimulus / regulation of immune response / immunoglobulin mediated immune response / positive regulation of phagocytosis / neutrophil chemotaxis / positive regulation of calcium-mediated signaling / SH2 domain binding / Neutrophil degranulation / osteoclast differentiation / integrin-mediated signaling pathway / establishment of localization in cell / protein localization to plasma membrane / phosphoprotein binding / calcium-mediated signaling / receptor internalization / positive regulation of interleukin-6 production / positive regulation of peptidyl-tyrosine phosphorylation / antigen processing and presentation of exogenous peptide antigen via MHC class II / positive regulation of tumor necrosis factor production / positive regulation of immune response / cell surface receptor signaling pathway / endosome / defense response to bacterium / immune response / protein heterodimerization activity / membrane raft / external side of plasma membrane / innate immune response / protein kinase binding / cell surface / signal transduction / protein homodimerization activity / identical protein binding / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.71 Å | |||||||||
 Authors Authors | Zhang Z / Yui M / Ohto U / Shimizu T | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Mouse Fc epsilon RI in complex withMouse Fc epsilon RI in complex with mIgE Fc mIgE Fc Authors: Zhang Z / Yui M / Ohto U / Shimizu T | |||||||||
| History |
|
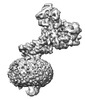
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36941.map.gz emd_36941.map.gz | 49.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36941-v30.xml emd-36941-v30.xml emd-36941.xml emd-36941.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36941.png emd_36941.png | 85 KB | ||
| Filedesc metadata |  emd-36941.cif.gz emd-36941.cif.gz | 6 KB | ||
| Others |  emd_36941_additional_1.map.gz emd_36941_additional_1.map.gz emd_36941_half_map_1.map.gz emd_36941_half_map_1.map.gz emd_36941_half_map_2.map.gz emd_36941_half_map_2.map.gz | 26.5 MB 49 MB 49 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36941 http://ftp.pdbj.org/pub/emdb/structures/EMD-36941 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36941 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36941 | HTTPS FTP |
-Validation report
| Summary document |  emd_36941_validation.pdf.gz emd_36941_validation.pdf.gz | 860.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36941_full_validation.pdf.gz emd_36941_full_validation.pdf.gz | 860.1 KB | Display | |
| Data in XML |  emd_36941_validation.xml.gz emd_36941_validation.xml.gz | 11.7 KB | Display | |
| Data in CIF |  emd_36941_validation.cif.gz emd_36941_validation.cif.gz | 13.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36941 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36941 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36941 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36941 | HTTPS FTP |
-Related structure data
| Related structure data |  8k7tMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36941.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36941.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.245 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_36941_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36941_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36941_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mouse Fc epsilon RI in complex with mIgE Fc
| Entire | Name: Mouse Fc epsilon RI in complex with mIgE Fc |
|---|---|
| Components |
|
-Supramolecule #1: Mouse Fc epsilon RI in complex with mIgE Fc
| Supramolecule | Name: Mouse Fc epsilon RI in complex with mIgE Fc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: High affinity immunoglobulin epsilon receptor subunit alpha
| Macromolecule | Name: High affinity immunoglobulin epsilon receptor subunit alpha type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.646197 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MVTGRSAQLC LALLFMSLDV ILTATEKSVL TLDPPWIRIF TGEKVTLSCY GNNHLQMNST TKWIHNGTVS EVNSSHLVIV SATVQDSGK YICQKQGLFK SKPVYLNVTQ DWLLLQTSAD MVLVHGSFDI RCHGWKNWNV RKVIYYRNDH AFNYSYESPV S IREATLND ...String: MVTGRSAQLC LALLFMSLDV ILTATEKSVL TLDPPWIRIF TGEKVTLSCY GNNHLQMNST TKWIHNGTVS EVNSSHLVIV SATVQDSGK YICQKQGLFK SKPVYLNVTQ DWLLLQTSAD MVLVHGSFDI RCHGWKNWNV RKVIYYRNDH AFNYSYESPV S IREATLND SGTYHCKGYL RQVKYESDKF RIAVVKAYKC KYYWLQLIFP LLVAILFAVD TGLLLSTEEQ FKSVLEIQKT GK YKKVETE LLTENLYFQG DYKDDDDKHH HHHHHH UniProtKB: High affinity immunoglobulin epsilon receptor subunit alpha |
-Macromolecule #2: High affinity immunoglobulin epsilon receptor subunit gamma
| Macromolecule | Name: High affinity immunoglobulin epsilon receptor subunit gamma type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.890856 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MISAVILFLL LLVEQAAALG EPQLCYILDA VLFLYGIVLT LLYCRLKIQV RKAAIASREK ADAVYTGLNT RSQETYETLK HEKPPQWSH PQFEKEQKLI SEEDL UniProtKB: High affinity immunoglobulin epsilon receptor subunit gamma |
-Macromolecule #3: High affinity immunoglobulin epsilon receptor subunit beta
| Macromolecule | Name: High affinity immunoglobulin epsilon receptor subunit beta type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.166141 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDTENRSRAD LALPNPQESS SAPDIELLEA SPAKAAPPKQ TWRTFLKKEL EFLGATQILV GLICLCFGTI VCSVLYVSDF DEEVLLLYK LGYPFWGAVL FVLSGFLSII SERKNTLYLV RGSLGANIVS SIAAGTGIAM LILNLTNNFA YMNNCKNVTE D DGCFVASF ...String: MDTENRSRAD LALPNPQESS SAPDIELLEA SPAKAAPPKQ TWRTFLKKEL EFLGATQILV GLICLCFGTI VCSVLYVSDF DEEVLLLYK LGYPFWGAVL FVLSGFLSII SERKNTLYLV RGSLGANIVS SIAAGTGIAM LILNLTNNFA YMNNCKNVTE D DGCFVASF TTELVLMMLF LTILAFCSAV LFTIYRIGQE LESKKVPDDR LYEELNVYSP IYSELEDKGE TSSPVDSEQK LI SEEDL UniProtKB: High affinity immunoglobulin epsilon receptor subunit beta |
-Macromolecule #4: IgE Fc
| Macromolecule | Name: IgE Fc / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.369723 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METGLRWLLL VAVLKGVQCV RPVNITDPTL ELLHSSCDPN AFHSTIQLYC FIYGHILNDV SVSWLMDDRE ITDTLAQTVL IKEEGKLAS TCSKLNITEQ QWMSESTFTC KVTSQGVDYL AHTRRCPDHE PRGVITYLIP PSPLDLYQNG APKLTCLVVD L ESEKNVNV ...String: METGLRWLLL VAVLKGVQCV RPVNITDPTL ELLHSSCDPN AFHSTIQLYC FIYGHILNDV SVSWLMDDRE ITDTLAQTVL IKEEGKLAS TCSKLNITEQ QWMSESTFTC KVTSQGVDYL AHTRRCPDHE PRGVITYLIP PSPLDLYQNG APKLTCLVVD L ESEKNVNV TWNQEKKTSV SASQWYTKHH NNATTSITSI LPVVAKDWIE GYGYQCIVDH PDFPKPIVRS ITKTPGQRSA PE VYVFPPP EEESEDKRTL TCLIQNFFPE DISVQWLGDG KLISNSQHST TTPLKSNGSN QGFFIFSRLE VAKTLWTQRK QFT CQVIHE ALQKPRKLEK TISTSLGNTS LRPSHHHHHH |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 7 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 61.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.71 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 91911 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)