[English] 日本語
 Yorodumi
Yorodumi- EMDB-36390: Cryo-EM structure of the human nucleosome lacking N-terminal regi... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human nucleosome lacking N-terminal region of H2A, H2B, H3, and H4 | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | nuclear protein / chromatin / GENE REGULATION-DNA COMPLEX | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of tumor necrosis factor-mediated signaling pathway / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere ...negative regulation of tumor necrosis factor-mediated signaling pathway / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / epigenetic regulation of gene expression / telomere organization / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Interleukin-7 signaling / RNA Polymerase I Promoter Opening / Inhibition of DNA recombination at telomere / Assembly of the ORC complex at the origin of replication / Meiotic synapsis / SUMOylation of chromatin organization proteins / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / SIRT1 negatively regulates rRNA expression / HCMV Late Events / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / innate immune response in mucosa / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / HDACs deacetylate histones / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / lipopolysaccharide binding / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / HDMs demethylate histones / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Negative Regulation of CDH1 Gene Transcription / G2/M DNA damage checkpoint / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / PKMTs methylate histone lysines / DNA Damage/Telomere Stress Induced Senescence / Pre-NOTCH Transcription and Translation / Meiotic recombination / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Metalloprotease DUBs / Transcriptional regulation of granulopoiesis / RMTs methylate histone arginines / HCMV Early Events / structural constituent of chromatin / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / heterochromatin formation / nucleosome assembly / E3 ubiquitin ligases ubiquitinate target proteins / antibacterial humoral response / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / HATs acetylate histones / Factors involved in megakaryocyte development and platelet production / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromatin organization / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / Processing of DNA double-strand break ends / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / killing of cells of another organism / defense response to Gram-negative bacterium / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / cadherin binding / Amyloid fiber formation / protein heterodimerization activity / negative regulation of cell population proliferation / protein-containing complex / : / DNA binding / RNA binding / extracellular exosome / extracellular region / nucleoplasm / membrane / nucleus / cytosol Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||||||||||||||
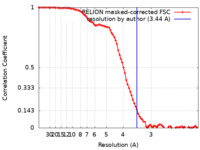
| Method | single particle reconstruction / cryo EM / Resolution: 3.44 Å | |||||||||||||||||||||
 Authors Authors | Oishi T / Hatazawa S / Kujirai T / Kato J / Kobayashi Y / Ogasawara M / Akatsu M / Takizawa Y / Kurumizaka H | |||||||||||||||||||||
| Funding support |  Japan, 6 items Japan, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Contributions of histone tail clipping and acetylation in nucleosome transcription by RNA polymerase II. Authors: Takumi Oishi / Suguru Hatazawa / Tomoya Kujirai / Junko Kato / Yuki Kobayashi / Mitsuo Ogasawara / Munetaka Akatsu / Haruhiko Ehara / Shun-Ichi Sekine / Gosuke Hayashi / Yoshimasa Takizawa / ...Authors: Takumi Oishi / Suguru Hatazawa / Tomoya Kujirai / Junko Kato / Yuki Kobayashi / Mitsuo Ogasawara / Munetaka Akatsu / Haruhiko Ehara / Shun-Ichi Sekine / Gosuke Hayashi / Yoshimasa Takizawa / Hitoshi Kurumizaka /  Abstract: The N-terminal tails of histones protrude from the nucleosome core and are target sites for histone modifications, such as acetylation and methylation. Histone acetylation is considered to enhance ...The N-terminal tails of histones protrude from the nucleosome core and are target sites for histone modifications, such as acetylation and methylation. Histone acetylation is considered to enhance transcription in chromatin. However, the contribution of the histone N-terminal tail to the nucleosome transcription by RNA polymerase II (RNAPII) has not been clarified. In the present study, we reconstituted nucleosomes lacking the N-terminal tail of each histone, H2A, H2B, H3 or H4, and performed RNAPII transcription assays. We found that the N-terminal tail of H3, but not H2A, H2B and H4, functions in RNAPII pausing at the SHL(-5) position of the nucleosome. Consistently, the RNAPII transcription assay also revealed that the nucleosome containing N-terminally acetylated H3 drastically alleviates RNAPII pausing at the SHL(-5) position. In addition, the H3 acetylated nucleosome produced increased amounts of the run-off transcript. These results provide important evidence that the H3 N-terminal tail plays a role in RNAPII pausing at the SHL(-5) position of the nucleosome, and its acetylation directly alleviates this nucleosome barrier. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36390.map.gz emd_36390.map.gz | 33.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36390-v30.xml emd-36390-v30.xml emd-36390.xml emd-36390.xml | 26.8 KB 26.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36390_fsc.xml emd_36390_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_36390.png emd_36390.png | 70 KB | ||
| Filedesc metadata |  emd-36390.cif.gz emd-36390.cif.gz | 6.7 KB | ||
| Others |  emd_36390_additional_1.map.gz emd_36390_additional_1.map.gz emd_36390_half_map_1.map.gz emd_36390_half_map_1.map.gz emd_36390_half_map_2.map.gz emd_36390_half_map_2.map.gz | 7.4 MB 40.8 MB 40.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36390 http://ftp.pdbj.org/pub/emdb/structures/EMD-36390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36390 | HTTPS FTP |
-Related structure data
| Related structure data |  8jlaMC  8jl9C  8jlbC  8jldC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36390.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36390.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_36390_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
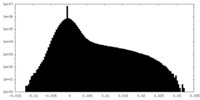
| Density Histograms |
-Half map: #1
| File | emd_36390_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36390_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The human nucleosome lacking N-terminal region of H2A, H2B, H3, a...
| Entire | Name: The human nucleosome lacking N-terminal region of H2A, H2B, H3, and H4 with scFv |
|---|---|
| Components |
|
-Supramolecule #1: The human nucleosome lacking N-terminal region of H2A, H2B, H3, a...
| Supramolecule | Name: The human nucleosome lacking N-terminal region of H2A, H2B, H3, and H4 with scFv type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Human nucleosome lacking N-terminal region of H2A, H2B, H3, and H4 with scFv |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Histone H3.1
| Macromolecule | Name: Histone H3.1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.816991 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMSAPATG GVKKPHRYRP GTVALREIRR YQKSTELLIR KLPFQRLVRE IAQDFKTDLR FQSSAVMALQ EACEAYLVGL FEDTNLCAI HAKRVTIMPK DIQLARRIRG ERA UniProtKB: Histone H3.1 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.404241 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMKRHRKV LRDNIQGITK PAIRRLARRG GVKRISGLIY EETRGVLKVF LENVIRDAVT YTEHAKRKTV TAMDVVYALK RQGRTLYGF GG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A type 1-B/E
| Macromolecule | Name: Histone H2A type 1-B/E / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.58886 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMARAKAK TRSSRAGLQF PVGRVHRLLR KGNYSERVGA GAPVYLAAVL EYLTAEILEL AGNAARDNKK TRIIPRHLQL AIRNDEELN KLLGRVTIAQ GGVLPNIQAV LLPKKTESHH KAKGK UniProtKB: Histone H2A type 1-B/E |
-Macromolecule #4: Histone H2B type 1-J
| Macromolecule | Name: Histone H2B type 1-J / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.751545 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSHMDGKKRK RSRKESYSIY VYKVLKQVHP DTGISSKAMG IMNSFVNDIF ERIAGEASRL AHYNKRSTIT SREIQTAVRL LLPGELAKH AVSEGTKAVT KYTSAK UniProtKB: Histone H2B type 1-J |
-Macromolecule #5: DNA (193-MER)
| Macromolecule | Name: DNA (193-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 59.589984 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DC)(DG)(DT)(DA)(DA)(DT) (DA)(DT)(DT)(DG)(DG)(DC)(DC)(DA)(DG)(DC) (DT)(DA)(DG)(DG)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG) (DC) (DC)(DG)(DA)(DG)(DG)(DC) ...String: (DA)(DT)(DC)(DA)(DC)(DG)(DT)(DA)(DA)(DT) (DA)(DT)(DT)(DG)(DG)(DC)(DC)(DA)(DG)(DC) (DT)(DA)(DG)(DG)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG) (DC) (DC)(DG)(DA)(DG)(DG)(DC)(DC)(DG) (DC)(DT)(DC)(DA)(DA)(DT)(DT)(DG)(DG)(DT) (DC)(DG) (DT)(DA)(DG)(DA)(DC)(DA)(DG) (DC)(DT)(DC)(DT)(DA)(DG)(DC)(DA)(DC)(DC) (DG)(DC)(DT) (DT)(DA)(DA)(DA)(DC)(DG) (DC)(DA)(DC)(DG)(DT)(DA)(DC)(DG)(DG)(DA) (DA)(DT)(DC)(DC) (DG)(DT)(DA)(DC)(DG) (DT)(DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG) (DC)(DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC) (DG)(DA)(DC)(DC)(DA)(DA) (DT)(DT)(DG) (DA)(DG)(DC)(DG)(DG)(DC)(DC)(DT)(DC)(DG) (DG)(DC)(DA)(DC)(DC)(DG)(DG) (DG)(DA) (DT)(DT)(DG)(DT)(DG)(DA)(DT)(DC)(DC)(DT) (DA)(DG)(DC)(DT)(DG)(DG)(DC)(DC) (DA) (DA)(DT)(DA)(DT)(DT)(DA)(DC)(DG)(DT)(DG) (DA)(DT) |
-Macromolecule #6: DNA (193-MER)
| Macromolecule | Name: DNA (193-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 59.580969 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DC)(DG)(DT)(DA)(DA)(DT) (DA)(DT)(DT)(DG)(DG)(DC)(DC)(DA)(DG)(DC) (DT)(DA)(DG)(DG)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG) (DC) (DC)(DG)(DA)(DG)(DG)(DC) ...String: (DA)(DT)(DC)(DA)(DC)(DG)(DT)(DA)(DA)(DT) (DA)(DT)(DT)(DG)(DG)(DC)(DC)(DA)(DG)(DC) (DT)(DA)(DG)(DG)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DC)(DG)(DG)(DT)(DG) (DC) (DC)(DG)(DA)(DG)(DG)(DC)(DC)(DG) (DC)(DT)(DC)(DA)(DA)(DT)(DT)(DG)(DG)(DT) (DC)(DG) (DT)(DA)(DG)(DA)(DC)(DA)(DG) (DC)(DT)(DC)(DT)(DA)(DG)(DC)(DA)(DC)(DC) (DG)(DC)(DT) (DT)(DA)(DA)(DA)(DC)(DG) (DC)(DA)(DC)(DG)(DT)(DA)(DC)(DG)(DG)(DA) (DT)(DT)(DC)(DC) (DG)(DT)(DA)(DC)(DG) (DT)(DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG) (DC)(DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC) (DG)(DA)(DC)(DC)(DA)(DA) (DT)(DT)(DG) (DA)(DG)(DC)(DG)(DG)(DC)(DC)(DT)(DC)(DG) (DG)(DC)(DA)(DC)(DC)(DG)(DG) (DG)(DA) (DT)(DT)(DG)(DT)(DG)(DA)(DT)(DC)(DC)(DT) (DA)(DG)(DC)(DT)(DG)(DG)(DC)(DC) (DA) (DA)(DT)(DA)(DT)(DT)(DA)(DC)(DG)(DT)(DG) (DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | Human nucleosome lacking N-terminal region of H2A, H2B, H3, and H4 aided by scFv |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 59.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)