+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of APOBEC3G-Vif complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Human antiviral protein / HIV / ANTIVIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationRUNX3 regulates RUNX1-mediated transcription / RUNX1 regulates transcription of genes involved in BCR signaling / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation / RUNX2 regulates chondrocyte maturation / negative regulation of CD4-positive, alpha-beta T cell differentiation / RUNX1 and FOXP3 control the development of regulatory T lymphocytes (Tregs) ...RUNX3 regulates RUNX1-mediated transcription / RUNX1 regulates transcription of genes involved in BCR signaling / RUNX1 regulates transcription of genes involved in interleukin signaling / RUNX2 regulates bone development / core-binding factor complex / RUNX1 regulates expression of components of tight junctions / positive regulation of CD8-positive, alpha-beta T cell differentiation / RUNX2 regulates chondrocyte maturation / negative regulation of CD4-positive, alpha-beta T cell differentiation / RUNX1 and FOXP3 control the development of regulatory T lymphocytes (Tregs) / lymphocyte differentiation / RUNX2 regulates genes involved in cell migration / Transcriptional regulation by RUNX2 / RUNX2 regulates genes involved in differentiation of myeloid cells / RUNX1 regulates transcription of genes involved in differentiation of keratinocytes / RUNX3 Regulates Immune Response and Cell Migration / myeloid cell differentiation / definitive hemopoiesis / RUNX1 regulates transcription of genes involved in differentiation of myeloid cells / Regulation of RUNX1 Expression and Activity / RUNX1 regulates transcription of genes involved in WNT signaling / RUNX1 regulates estrogen receptor mediated transcription / RUNX2 regulates osteoblast differentiation / RUNX1 interacts with co-factors whose precise effect on RUNX1 targets is not known / RUNX3 regulates p14-ARF / cell maturation / Regulation of RUNX3 expression and activity / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Transcriptional regulation of granulopoiesis / protein polyubiquitination / Regulation of RUNX2 expression and activity / osteoblast differentiation / RUNX1 regulates transcription of genes involved in differentiation of HSCs / transcription by RNA polymerase II / Estrogen-dependent gene expression / sequence-specific DNA binding / transcription coactivator activity / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / nucleoplasm / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human immunodeficiency virus 1 / synthetic construct (others) Human immunodeficiency virus 1 / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Kouno T / Shibata S / Hyun J / Kim TG / Wolf M | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural insights into RNA bridging between HIV-1 Vif and antiviral factor APOBEC3G. Authors: Takahide Kouno / Satoshi Shibata / Megumi Shigematsu / Jaekyung Hyun / Tae Gyun Kim / Hiroshi Matsuo / Matthias Wolf /     Abstract: Great effort has been devoted to discovering the basis of A3G-Vif interaction, the key event of HIV's counteraction mechanism to evade antiviral innate immune response. Here we show reconstitution of ...Great effort has been devoted to discovering the basis of A3G-Vif interaction, the key event of HIV's counteraction mechanism to evade antiviral innate immune response. Here we show reconstitution of the A3G-Vif complex and subsequent A3G ubiquitination in vitro and report the cryo-EM structure of the A3G-Vif complex at 2.8 Å resolution using solubility-enhanced variants of A3G and Vif. We present an atomic model of the A3G-Vif interface, which assembles via known amino acid determinants. This assembly is not achieved by protein-protein interaction alone, but also involves RNA. The cryo-EM structure and in vitro ubiquitination assays identify an adenine/guanine base preference for the interaction and a unique Vif-ribose contact. This establishes the biological significance of an RNA ligand. Further assessment of interactions between A3G, Vif, and RNA ligands show that the A3G-Vif assembly and subsequent ubiquitination can be controlled by amino acid mutations at the interface or by polynucleotide modification, suggesting that a specific chemical moiety would be a promising pharmacophore to inhibit the A3G-Vif interaction. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35999.map.gz emd_35999.map.gz | 169.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35999-v30.xml emd-35999-v30.xml emd-35999.xml emd-35999.xml | 20.6 KB 20.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35999.png emd_35999.png | 117.9 KB | ||
| Filedesc metadata |  emd-35999.cif.gz emd-35999.cif.gz | 6.6 KB | ||
| Others |  emd_35999_half_map_1.map.gz emd_35999_half_map_1.map.gz emd_35999_half_map_2.map.gz emd_35999_half_map_2.map.gz | 169.8 MB 170 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35999 http://ftp.pdbj.org/pub/emdb/structures/EMD-35999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35999 | HTTPS FTP |
-Related structure data
| Related structure data |  8j62MC  8h0iC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35999.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35999.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.5696 Å | ||||||||||||||||||||||||||||||||||||
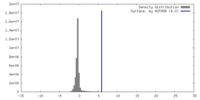
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35999_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
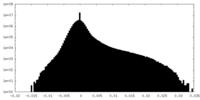
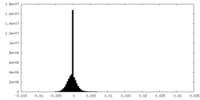
| Density Histograms |
-Half map: #1
| File | emd_35999_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
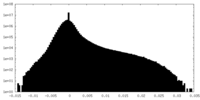
| Density Histograms |
- Sample components
Sample components
-Entire : A3G-VC-RNA complex
| Entire | Name: A3G-VC-RNA complex |
|---|---|
| Components |
|
-Supramolecule #1: A3G-VC-RNA complex
| Supramolecule | Name: A3G-VC-RNA complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: APOBEC3G,COre binding factor
| Supramolecule | Name: APOBEC3G,COre binding factor / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1, #3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Viral infectibity factor
| Supramolecule | Name: Viral infectibity factor / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #4: RNA
| Supramolecule | Name: RNA / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|
-Macromolecule #1: APOBEC3G
| Macromolecule | Name: APOBEC3G / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.857266 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGHMDPDTF SYNFNNRPIL SRRNTVWLCY EVERLDNGTW VKMDQHRGQV YSELKYHPEM RFLSLVSKWK LHRDQEYEVT WYISWSPCT KCARDMATFL QENTHVTLTI FVARLYYFWD PDYQEALRSL AQAGATIKIM NYDEFQHCWS KFVYSQGAPF Q PWDGLDEY ...String: GPGHMDPDTF SYNFNNRPIL SRRNTVWLCY EVERLDNGTW VKMDQHRGQV YSELKYHPEM RFLSLVSKWK LHRDQEYEVT WYISWSPCT KCARDMATFL QENTHVTLTI FVARLYYFWD PDYQEALRSL AQAGATIKIM NYDEFQHCWS KFVYSQGAPF Q PWDGLDEY SQALSGMLGE ILRHSMDPPT FTFNFNNEPW VRGRHETYLC YEVERMHNDT WVKLNQRRGF LANQAPHKHG FL EGRHAEL CFLDVIPFWK LDLDQDYRVT CFTSWSPCFS CAQEMAKFIS KNKHVSLCIK TARIYDDQGR AQEGLRTLAE AGA KISIMT YSEFKHCWDT FVDHQGAPFQ PWDGLDEHSQ DLSGRLRAIL QNQEN |
-Macromolecule #2: Viral infectivity factor
| Macromolecule | Name: Viral infectivity factor / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 18.180643 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGHHHHHHSQ DPMENRWQVM IVWQVDRMRI NTWKRLVKHH MYISRKAKDW FYRHHYESTN PKISSEVHIP LGDAKLVITT YWGLHTGER DWHLGQGVSI EWRKKRYSTQ VDPDLADQLI HLHYFDEASE GSQIKPPLPS VRKLTEDRWN K |
-Macromolecule #3: Core binding factor beta
| Macromolecule | Name: Core binding factor beta / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.528727 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPRVVPDQRS KFENEEFFRK LSRECEIKYT GFRDRPHEER QARFQNACRD GRSEIAFVAT GTNLSLQFFP ASWQGEQRQT PSREYVDLE REAGKVYLKA PMILNGVCVI WKGWIDLQRL DGMGCLEFDE ERAQQEDALA QQAFEEARRR TREFEDR UniProtKB: Core-binding factor subunit beta |
-Macromolecule #4: RNA (5'-R(*CP*GP*GP*UP*UP*GP*AP*UP*UP*GP*UP*UP*UP*UP*AP*AP*CP*AP*...
| Macromolecule | Name: RNA (5'-R(*CP*GP*GP*UP*UP*GP*AP*UP*UP*GP*UP*UP*UP*UP*AP*AP*CP*AP*A)-3') type: rna / ID: 4 / Number of copies: 2 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 6.370791 KDa |
| Sequence | String: CGGUUGAUUG UUUUAACAAA |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Sugar embedding | Material: Graphene oxide | ||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: PLASMA CLEANING | ||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 37.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 2.5 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1.3) / Number images used: 186943 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1.3) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1.3) |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)