+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | top segment of the bacteriophage M13 mini variant | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral coat protein / M13 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhelical viral capsid / host cell membrane / virion component / host cell plasma membrane / membrane Similarity search - Function | |||||||||
| Biological species |  Inovirus M13 Inovirus M13 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Xiang Y / Jia Q | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure of a bacteriophage M13 mini variant. Authors: Qi Jia / Ye Xiang /  Abstract: Filamentous bacteriophages package their circular, single stranded DNA genome with the major coat protein pVIII and the minor coat proteins pIII, pVII, pVI, and pIX. Here, we report the cryo-EM ...Filamentous bacteriophages package their circular, single stranded DNA genome with the major coat protein pVIII and the minor coat proteins pIII, pVII, pVI, and pIX. Here, we report the cryo-EM structure of a ~500 Å long bacteriophage M13 mini variant. The distal ends of the mini phage are sealed by two cap-like complexes composed of the minor coat proteins. The top cap complex consists of pVII and pIX, both exhibiting a single helix structure. Arg33 of pVII and Glu29 of pIX, located on the inner surface of the cap, play a key role in recognizing the genome packaging signal. The bottom cap complex is formed by the hook-like structures of pIII and pVI, arranged in helix barrels. Most of the inner ssDNA genome adopts a double helix structure with a similar pitch to that of the A-form double-stranded DNA. These findings provide insights into the assembly of filamentous bacteriophages. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35795.map.gz emd_35795.map.gz | 5.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35795-v30.xml emd-35795-v30.xml emd-35795.xml emd-35795.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
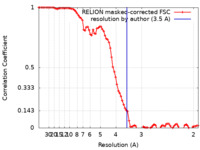
| FSC (resolution estimation) |  emd_35795_fsc.xml emd_35795_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_35795.png emd_35795.png | 31.4 KB | ||
| Others |  emd_35795_half_map_1.map.gz emd_35795_half_map_1.map.gz emd_35795_half_map_2.map.gz emd_35795_half_map_2.map.gz | 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35795 http://ftp.pdbj.org/pub/emdb/structures/EMD-35795 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35795 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35795 | HTTPS FTP |
-Related structure data
| Related structure data |  8ixlMC  8jwwM  8ixjC  8ixkC  8jwtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_35795.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35795.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.97 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35795_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
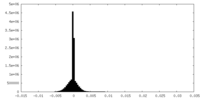
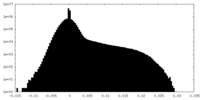
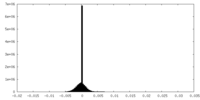
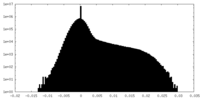
| Density Histograms |
-Half map: #1
| File | emd_35795_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : top segment of the bacteriophage M13 mini variant
| Entire | Name: top segment of the bacteriophage M13 mini variant |
|---|---|
| Components |
|
-Supramolecule #1: top segment of the bacteriophage M13 mini variant
| Supramolecule | Name: top segment of the bacteriophage M13 mini variant / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
-Macromolecule #1: Capsid protein G8P
| Macromolecule | Name: Capsid protein G8P / type: protein_or_peptide / ID: 1 / Number of copies: 25 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 5.243014 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AEGDDPAKAA FNSLQASATE YIGYAWAMVV VIVGATIGIK LFKKFTSKAS UniProtKB: Capsid protein G8P |
-Macromolecule #2: Tail virion protein G9P
| Macromolecule | Name: Tail virion protein G9P / type: protein_or_peptide / ID: 2 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 3.65527 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSVLVYSFAS FVLGWCLRSG ITYFTRLMET SS UniProtKB: Tail virion protein G9P |
-Macromolecule #3: Tail virion protein G7P
| Macromolecule | Name: Tail virion protein G7P / type: protein_or_peptide / ID: 3 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 3.603215 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEQVADFDTI YQAMIQISVV LCFALGIIAG GQR UniProtKB: Tail virion protein G7P |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)