+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | bottom segment of the bacteriophage M13 mini variant | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral coat protein / M13 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationviral extrusion / virion attachment to host cell pilus / adhesion receptor-mediated virion attachment to host cell / helical viral capsid / host cell membrane / virion component / viral capsid / entry receptor-mediated virion attachment to host cell / symbiont entry into host cell / host cell plasma membrane / membrane Similarity search - Function | |||||||||
| Biological species |  Inovirus M13 Inovirus M13 | |||||||||
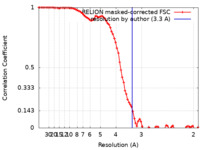
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Xiang Y / Jia Q | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure of a bacteriophage M13 mini variant. Authors: Qi Jia / Ye Xiang /  Abstract: Filamentous bacteriophages package their circular, single stranded DNA genome with the major coat protein pVIII and the minor coat proteins pIII, pVII, pVI, and pIX. Here, we report the cryo-EM ...Filamentous bacteriophages package their circular, single stranded DNA genome with the major coat protein pVIII and the minor coat proteins pIII, pVII, pVI, and pIX. Here, we report the cryo-EM structure of a ~500 Å long bacteriophage M13 mini variant. The distal ends of the mini phage are sealed by two cap-like complexes composed of the minor coat proteins. The top cap complex consists of pVII and pIX, both exhibiting a single helix structure. Arg33 of pVII and Glu29 of pIX, located on the inner surface of the cap, play a key role in recognizing the genome packaging signal. The bottom cap complex is formed by the hook-like structures of pIII and pVI, arranged in helix barrels. Most of the inner ssDNA genome adopts a double helix structure with a similar pitch to that of the A-form double-stranded DNA. These findings provide insights into the assembly of filamentous bacteriophages. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35794.map.gz emd_35794.map.gz | 6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35794-v30.xml emd-35794-v30.xml emd-35794.xml emd-35794.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35794_fsc.xml emd_35794_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_35794.png emd_35794.png | 42.1 KB | ||
| Others |  emd_35794_half_map_1.map.gz emd_35794_half_map_1.map.gz emd_35794_half_map_2.map.gz emd_35794_half_map_2.map.gz | 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35794 http://ftp.pdbj.org/pub/emdb/structures/EMD-35794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35794 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35794 | HTTPS FTP |
-Validation report
| Summary document |  emd_35794_validation.pdf.gz emd_35794_validation.pdf.gz | 757.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35794_full_validation.pdf.gz emd_35794_full_validation.pdf.gz | 757 KB | Display | |
| Data in XML |  emd_35794_validation.xml.gz emd_35794_validation.xml.gz | 14.3 KB | Display | |
| Data in CIF |  emd_35794_validation.cif.gz emd_35794_validation.cif.gz | 20 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35794 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35794 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35794 | HTTPS FTP |
-Related structure data
| Related structure data |  8ixkMC  8jwxM  8ixjC  8ixlC  8jwtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_35794.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35794.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.97 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35794_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
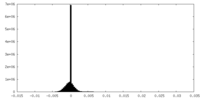
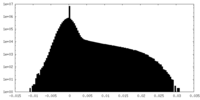
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35794_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
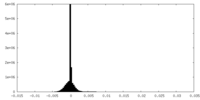
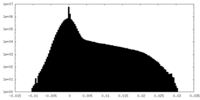
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : bottom segment of the bacteriophage M13 mini variant
| Entire | Name: bottom segment of the bacteriophage M13 mini variant |
|---|---|
| Components |
|
-Supramolecule #1: bottom segment of the bacteriophage M13 mini variant
| Supramolecule | Name: bottom segment of the bacteriophage M13 mini variant / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
-Macromolecule #1: Attachment protein G3P
| Macromolecule | Name: Attachment protein G3P / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 42.573766 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AETVESCLAK PHTENSFTNV WKDDKTLDRY ANYEGCLWNA TGVVVCTGDE TQCYGTWVPI GLAIPENEGG GSEGGGSEGG GSEGGGTKP PEYGDTPIPG YTYINPLDGT YPPGTEQNPA NPNPSLEESQ PLNTFMFQNN RFRNRQGALT VYTGTVTQGT D PVKTYYQY ...String: AETVESCLAK PHTENSFTNV WKDDKTLDRY ANYEGCLWNA TGVVVCTGDE TQCYGTWVPI GLAIPENEGG GSEGGGSEGG GSEGGGTKP PEYGDTPIPG YTYINPLDGT YPPGTEQNPA NPNPSLEESQ PLNTFMFQNN RFRNRQGALT VYTGTVTQGT D PVKTYYQY TPVSSKAMYD AYWNGKFRDC AFHSGFNEDP FVCEYQGQSS DLPQPPVNAG GGSGGGSGGG SEGGGSEGGG SE GGGSEGG GSGGGSGSGD FDYEKMANAN KGAMTENADE NALQSDAKGK LDSVATDYGA AIDGFIGDVS GLANGNGATG DFA GSNSQM AQVGDGDNSP LMNNFRQYLP SLPQSVECRP FVFGAGKPYE FSIDCDKINL FRGVFAFLLY VATFMYVFST FANI LRNKE S UniProtKB: Attachment protein G3P |
-Macromolecule #2: Capsid protein G8P
| Macromolecule | Name: Capsid protein G8P / type: protein_or_peptide / ID: 2 / Number of copies: 15 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 7.633036 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKSLVLKAS VAVATLVPML SFAAEGDDPA KAAFNSLQAS ATEYIGYAWA MVVVIVGATI GIKLFKKFTS KAS UniProtKB: Capsid protein G8P |
-Macromolecule #3: Head virion protein G6P
| Macromolecule | Name: Head virion protein G6P / type: protein_or_peptide / ID: 3 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Inovirus M13 Inovirus M13 |
| Molecular weight | Theoretical: 12.357984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPVLLGIPLL LRFLGFLLVT LFGYLLTFLK KGFGKIAIAI SLFLALIIGL NSILVGYLSD ISAQLPSDFV QGVQLILPSN ALPCFYVIL SVKAAIFIFD VKQKIVSYLD WDK UniProtKB: Head virion protein G6P |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.7 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X