[English] 日本語
 Yorodumi
Yorodumi- EMDB-34504: cryo-EM structure of cellodextrin phosphorylase from Clostridium ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of cellodextrin phosphorylase from Clostridium thermocellum | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cellulose / cellodextrin / phosphorolysis / synthesis / CARBOHYDRATE | |||||||||
| Function / homology |  Function and homology information Function and homology informationglycosyltransferase activity / carbohydrate binding / carbohydrate metabolic process Similarity search - Function | |||||||||
| Biological species |  Acetivibrio thermocellus (bacteria) Acetivibrio thermocellus (bacteria) | |||||||||
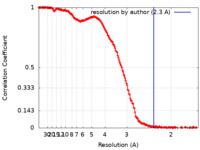
| Method | single particle reconstruction / cryo EM / Resolution: 2.3 Å | |||||||||
 Authors Authors | Kuga T / Sunagawa N / Igarashi K | |||||||||
| Funding support |  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: structure and dynamics of cellodextrin phosphorylase from Clostridium thermocellum determine chain length and crystalline packing of highly ordered cellulose II synthesized in vitro Authors: Kuga T / Sunagawa N / Igarashi K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34504.map.gz emd_34504.map.gz | 217.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34504-v30.xml emd-34504-v30.xml emd-34504.xml emd-34504.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34504_fsc.xml emd_34504_fsc.xml | 21.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_34504.png emd_34504.png | 78.5 KB | ||
| Filedesc metadata |  emd-34504.cif.gz emd-34504.cif.gz | 6.2 KB | ||
| Others |  emd_34504_half_map_1.map.gz emd_34504_half_map_1.map.gz emd_34504_half_map_2.map.gz emd_34504_half_map_2.map.gz | 391.7 MB 391.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34504 http://ftp.pdbj.org/pub/emdb/structures/EMD-34504 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34504 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34504 | HTTPS FTP |
-Validation report
| Summary document |  emd_34504_validation.pdf.gz emd_34504_validation.pdf.gz | 800.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34504_full_validation.pdf.gz emd_34504_full_validation.pdf.gz | 800 KB | Display | |
| Data in XML |  emd_34504_validation.xml.gz emd_34504_validation.xml.gz | 24.1 KB | Display | |
| Data in CIF |  emd_34504_validation.cif.gz emd_34504_validation.cif.gz | 31.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34504 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34504 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34504 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34504 | HTTPS FTP |
-Related structure data
| Related structure data |  8h6hMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34504.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34504.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34504_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
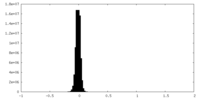
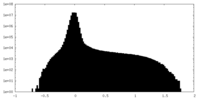
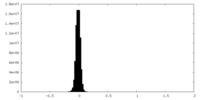
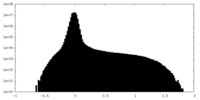
| Density Histograms |
-Half map: #1
| File | emd_34504_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimeric structure of cellodextrin phosphorylase from Clostrid...
| Entire | Name: Homodimeric structure of cellodextrin phosphorylase from Clostridium thermocellum (CtCDP) |
|---|---|
| Components |
|
-Supramolecule #1: Homodimeric structure of cellodextrin phosphorylase from Clostrid...
| Supramolecule | Name: Homodimeric structure of cellodextrin phosphorylase from Clostridium thermocellum (CtCDP) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Acetivibrio thermocellus (bacteria) / Strain: YM4 Acetivibrio thermocellus (bacteria) / Strain: YM4 |
| Molecular weight | Theoretical: 240 KDa |
-Macromolecule #1: Cellodextrin phosphorylase
| Macromolecule | Name: Cellodextrin phosphorylase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acetivibrio thermocellus (bacteria) Acetivibrio thermocellus (bacteria) |
| Molecular weight | Theoretical: 112.599523 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVTKVTARNN KITPVELLNQ KFGNKINLGN FADAVFTDAA FKNVAGIANL PMKAPVMQVL MENCIVSKYL KQFVPDRSVC FVEEGQKFY IVLEDGQKIE VPEDVNKALK ATVSDVKHWA GYLTEDGEHV IDLLKPAPGP HFYVNLLIGN RLGFKRTLQT T PKSVVDRF ...String: MVTKVTARNN KITPVELLNQ KFGNKINLGN FADAVFTDAA FKNVAGIANL PMKAPVMQVL MENCIVSKYL KQFVPDRSVC FVEEGQKFY IVLEDGQKIE VPEDVNKALK ATVSDVKHWA GYLTEDGEHV IDLLKPAPGP HFYVNLLIGN RLGFKRTLQT T PKSVVDRF GRGSFRSHAA TQVLATRFDM RQEENGFPAN RQFYLYEDGK QIFYSALIDD NIVEATCKHS CNRTVIKYKT AC NLEITRT IFLVPHKKGF PLATELQRIE IKNASDKARN LSITYTGMFG TGAVHAIFED VTYTNVIMQS AALYNDKGEF IGI TPDYYP EEFKQDTRFV TMIVRNGDEK SFPQSFCTDY NDFVGTGTLE HPAGGCNLNN KLNRKGPGFF ALGAPFTVEP GKTV IIDTF TGLSSSKDNE NYSDAVMLRE LDNLLRYFEK SESVEETLNE IINFHENYGK YFQFNTGNKL FDSGFNRNLA FQVLY QTFM SRSFGQTQKG YREIGFREIQ DLFASMYYFI NIGYQDFVKE LLFEWTANVY KMGYANHNFY WVGKQPGLYS DDSLWL LQA YYRYIIYTKD TSVLNEEVPV ADGNNEKRAV RETLKAIIQY SACISVGDHG LPLLDLADWN DCLKIDSNSI DGATKEK LY YEQLKKTNGK YGDRFMSDYS ESVMNAFLLK LAIDHLAEIA TLDNDTQLAQ QMSELSKEVT DRIQKHAWKE NFFARVLI N RYKDGSYTYL GAKGDKLSAD PNIDGVYFLN SFAWSVLSDV ATDEQIAIMV DVIKKHLLTP YGLRLVTPAD LNKIANDTA TGHYFFGDRE NGAVFKHASM MAVAALIKAA KKVKDNELAK EMARIAYFMI DLVLPYKNLE NPFQVAGNPR ICTQYINTDT GENIGPLLS GTATWLNLNL ISLAGIEYTR DGISFNPILR EEETQLNFTL KAPKCSYKFS ITKPVGFARM ESSEYELFVD G QKIDNTVI PMYTDEKEHI VTLKFKHHHH HH UniProtKB: Cellodextrin phosphorylase |
-Macromolecule #2: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.0 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 20 mM Tris-HCl, 120 mM NaCl, 0.1 mM DTT | ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 280 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 49.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X