+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of a bacterial dioxygenase | |||||||||
 Map data Map data | Cryo-EM structure of a bacterial dioxygenase | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | heterohexamer / dioxygenase / FLAVOPROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information3-phenylpropionate catabolic process / dioxygenase activity / 2 iron, 2 sulfur cluster binding / oxidoreductase activity / iron ion binding Similarity search - Function | |||||||||
| Biological species |  Variovorax paradoxus (bacteria) Variovorax paradoxus (bacteria) | |||||||||
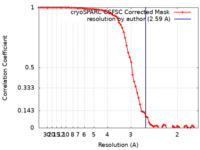
| Method | single particle reconstruction / cryo EM / Resolution: 2.59 Å | |||||||||
 Authors Authors | Yu G / Li Z / Zhang H | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2023 Journal: PLoS Biol / Year: 2023Title: Structural and biochemical characterization of the key components of an auxin degradation operon from the rhizosphere bacterium Variovorax. Authors: Yongjian Ma / Xuzichao Li / Feng Wang / Lingling Zhang / Shengmin Zhou / Xing Che / Dehao Yu / Xiang Liu / Zhuang Li / Huabing Sun / Guimei Yu / Heng Zhang /   Abstract: Plant-associated bacteria play important regulatory roles in modulating plant hormone auxin levels, affecting the growth and yields of crops. A conserved auxin degradation (iad) operon was recently ...Plant-associated bacteria play important regulatory roles in modulating plant hormone auxin levels, affecting the growth and yields of crops. A conserved auxin degradation (iad) operon was recently identified in the Variovorax genomes, which is responsible for root growth inhibition (RGI) reversion, promoting rhizosphere colonization and root growth. However, the molecular mechanism underlying auxin degradation by Variovorax remains unclear. Here, we systematically screened Variovorax iad operon products and identified 2 proteins, IadK2 and IadD, that directly associate with auxin indole-3-acetic acid (IAA). Further biochemical and structural studies revealed that IadK2 is a highly IAA-specific ATP-binding cassette (ABC) transporter solute-binding protein (SBP), likely involved in IAA uptake. IadD interacts with IadE to form a functional Rieske non-heme dioxygenase, which works in concert with a FMN-type reductase encoded by gene iadC to transform IAA into the biologically inactive 2-oxindole-3-acetic acid (oxIAA), representing a new bacterial pathway for IAA inactivation/degradation. Importantly, incorporation of a minimum set of iadC/D/E genes could enable IAA transformation by Escherichia coli, suggesting a promising strategy for repurposing the iad operon for IAA regulation. Together, our study identifies the key components and underlying mechanisms involved in IAA transformation by Variovorax and brings new insights into the bacterial turnover of plant hormones, which would provide the basis for potential applications in rhizosphere optimization and ecological agriculture. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34443.map.gz emd_34443.map.gz | 36.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34443-v30.xml emd-34443-v30.xml emd-34443.xml emd-34443.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34443_fsc.xml emd_34443_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_34443.png emd_34443.png | 103.8 KB | ||
| Filedesc metadata |  emd-34443.cif.gz emd-34443.cif.gz | 6.4 KB | ||
| Others |  emd_34443_half_map_1.map.gz emd_34443_half_map_1.map.gz emd_34443_half_map_2.map.gz emd_34443_half_map_2.map.gz | 37.7 MB 37.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34443 http://ftp.pdbj.org/pub/emdb/structures/EMD-34443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34443 | HTTPS FTP |
-Related structure data
| Related structure data |  8h2tMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34443.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34443.map.gz / Format: CCP4 / Size: 40.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of a bacterial dioxygenase | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_34443_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
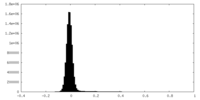
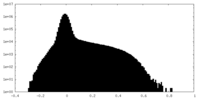
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_34443_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
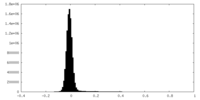
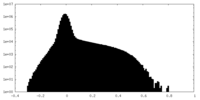
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : enzyme complex
| Entire | Name: enzyme complex |
|---|---|
| Components |
|
-Supramolecule #1: enzyme complex
| Supramolecule | Name: enzyme complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Variovorax paradoxus (bacteria) Variovorax paradoxus (bacteria) |
-Macromolecule #1: Rieske (2Fe-2S) domain protein
| Macromolecule | Name: Rieske (2Fe-2S) domain protein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Variovorax paradoxus (bacteria) / Strain: S110 Variovorax paradoxus (bacteria) / Strain: S110 |
| Molecular weight | Theoretical: 49.656898 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTSYRDNPDA IRALVQDDRV HRDLYTSQEL FELEQEHFFA NTWNYVGHES QLPKPGDWIS NEIAGRPLIV ARHSDGSVRA MMNRCAHKG SRLVNGPCGN TGKFFRCPYH AWTFKTDGSL LAIPLKTGYE NTALHECESA KGLTTLRYVR SHRGFIFVKI S DAGPDFDD ...String: MTSYRDNPDA IRALVQDDRV HRDLYTSQEL FELEQEHFFA NTWNYVGHES QLPKPGDWIS NEIAGRPLIV ARHSDGSVRA MMNRCAHKG SRLVNGPCGN TGKFFRCPYH AWTFKTDGSL LAIPLKTGYE NTALHECESA KGLTTLRYVR SHRGFIFVKI S DAGPDFDD YFGDSLSSID NMADRSPEGE LEIAGGCLRF MHQCNWKMFV ENLNDTMHPM VAHESSAGTA KRMWADKPED EP KPMAVEQ FAPFMSDYKF FEDMGIRTYD NGHSFTGVHF SIHSKYKAIP AYDDAMKARY GEAKTAQILG MARHNTVYYP NLT IKGAIQ AIRVVKPISA DRTLIESWTF RLKGAPPELL QRTTMYNRLI NSPFSVVGHD DLQAYRGMQA GLHASGNEWV SLHR NYDPS ELKGGEITTG GTNELPMRNQ YRAWVQRMTE TM UniProtKB: Rieske (2Fe-2S) domain protein |
-Macromolecule #2: Aromatic-ring-hydroxylating dioxygenase beta subunit
| Macromolecule | Name: Aromatic-ring-hydroxylating dioxygenase beta subunit / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Variovorax paradoxus (bacteria) Variovorax paradoxus (bacteria) |
| Molecular weight | Theoretical: 18.952352 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGTEVTRQD LIDFVVNEAH LLDTRRYEEW NALFTDDAFY WVPLVPDQED GLNHTSHLYE DKLLRELRIE RLKSPRAFSQ QPPSRCHHL LQVPVVEQFD AEGNRFVLRT GFHYTESQGD ELQFYVGTFF HHLTVRDGAL RMTLKRVNLL NCDAALPAVQ L FI UniProtKB: Aromatic-ring-hydroxylating dioxygenase beta subunit |
-Macromolecule #3: FE2/S2 (INORGANIC) CLUSTER
| Macromolecule | Name: FE2/S2 (INORGANIC) CLUSTER / type: ligand / ID: 3 / Number of copies: 3 / Formula: FES |
|---|---|
| Molecular weight | Theoretical: 175.82 Da |
| Chemical component information |  ChemComp-FES: |
-Macromolecule #4: 1H-INDOL-3-YLACETIC ACID
| Macromolecule | Name: 1H-INDOL-3-YLACETIC ACID / type: ligand / ID: 4 / Number of copies: 3 / Formula: IAC |
|---|---|
| Molecular weight | Theoretical: 175.184 Da |
| Chemical component information | 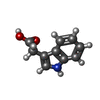 ChemComp-IAC: |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 3 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
| Details | No further treatment. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
 Movie
Movie Controller
Controller







 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)