+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | AK-42 inhibitor binding human ClC-2 TMD | |||||||||
 Map data Map data | CLC-2_AK42_TMD_3.5A | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | homo-dimer / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of aldosterone biosynthetic process / cell differentiation involved in salivary gland development / regulation of membrane depolarization during action potential / volume-sensitive chloride channel activity / astrocyte end-foot / stabilization of membrane potential / acinar cell differentiation / cellular hypotonic response / regulation of resting membrane potential / chloride channel regulator activity ...regulation of aldosterone biosynthetic process / cell differentiation involved in salivary gland development / regulation of membrane depolarization during action potential / volume-sensitive chloride channel activity / astrocyte end-foot / stabilization of membrane potential / acinar cell differentiation / cellular hypotonic response / regulation of resting membrane potential / chloride channel regulator activity / voltage-gated chloride channel activity / axon initial segment / dendritic spine membrane / chloride transport / phagocytosis, engulfment / positive regulation of oligodendrocyte differentiation / chloride channel complex / lung development / Stimuli-sensing channels / myelin sheath / retina development in camera-type eye / basolateral plasma membrane / perikaryon / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Wang L | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structures of ClC-2 chloride channel reveal the blocking mechanism of its specific inhibitor AK-42. Authors: Tao Ma / Lei Wang / Anping Chai / Chao Liu / Wenqiang Cui / Shuguang Yuan / Shannon Wing Ngor Au / Liang Sun / Xiaokang Zhang / Zhenzhen Zhang / Jianping Lu / Yuanzhu Gao / Peiyi Wang / ...Authors: Tao Ma / Lei Wang / Anping Chai / Chao Liu / Wenqiang Cui / Shuguang Yuan / Shannon Wing Ngor Au / Liang Sun / Xiaokang Zhang / Zhenzhen Zhang / Jianping Lu / Yuanzhu Gao / Peiyi Wang / Zhifang Li / Yujie Liang / Horst Vogel / Yu Tian Wang / Daping Wang / Kaige Yan / Huawei Zhang /   Abstract: ClC-2 transports chloride ions across plasma membranes and plays critical roles in cellular homeostasis. Its dysfunction is involved in diseases including leukodystrophy and primary aldosteronism. AK- ...ClC-2 transports chloride ions across plasma membranes and plays critical roles in cellular homeostasis. Its dysfunction is involved in diseases including leukodystrophy and primary aldosteronism. AK-42 was recently reported as a specific inhibitor of ClC-2. However, experimental structures are still missing to decipher its inhibition mechanism. Here, we present cryo-EM structures of apo ClC-2 and its complex with AK-42, both at 3.5 Å resolution. Residues S162, E205 and Y553 are involved in chloride binding and contribute to the ion selectivity. The side-chain of the gating glutamate E205 occupies the putative central chloride-binding site, indicating that our structure represents a closed state. Structural analysis, molecular dynamics and electrophysiological recordings identify key residues to interact with AK-42. Several AK-42 interacting residues are present in ClC-2 but not in other ClCs, providing a possible explanation for AK-42 specificity. Taken together, our results experimentally reveal the potential inhibition mechanism of ClC-2 inhibitor AK-42. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34202.map.gz emd_34202.map.gz | 4.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34202-v30.xml emd-34202-v30.xml emd-34202.xml emd-34202.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
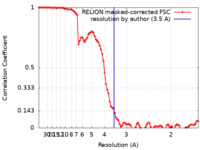
| FSC (resolution estimation) |  emd_34202_fsc.xml emd_34202_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_34202.png emd_34202.png | 39.1 KB | ||
| Masks |  emd_34202_msk_1.map emd_34202_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-34202.cif.gz emd-34202.cif.gz | 5.9 KB | ||
| Others |  emd_34202_half_map_1.map.gz emd_34202_half_map_1.map.gz emd_34202_half_map_2.map.gz emd_34202_half_map_2.map.gz | 48.6 MB 48.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34202 http://ftp.pdbj.org/pub/emdb/structures/EMD-34202 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34202 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34202 | HTTPS FTP |
-Related structure data
| Related structure data |  8gquMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34202.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34202.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC-2_AK42_TMD_3.5A | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_34202_msk_1.map emd_34202_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map
| File | emd_34202_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map
| File | emd_34202_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : AK-42 inhibitor binding human ClC-2 TMD 3.5A
| Entire | Name: AK-42 inhibitor binding human ClC-2 TMD 3.5A |
|---|---|
| Components |
|
-Supramolecule #1: AK-42 inhibitor binding human ClC-2 TMD 3.5A
| Supramolecule | Name: AK-42 inhibitor binding human ClC-2 TMD 3.5A / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 180 kDa/nm |
-Macromolecule #1: Chloride channel protein 2
| Macromolecule | Name: Chloride channel protein 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 98.642352 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAAAAAEEGM EPRALQYEQT LMYGRYTQDL GAFAKEEAAR IRLGGPEPWK GPPSSRAAPE LLEYGRSRCA RCRVCSVRCH KFLVSRVGE DWIFLVLLGL LMALVSWVMD YAIAACLQAQ QWMSRGLNTS ILLQYLAWVT YPVVLITFSA GFTQILAPQA V GSGIPEMK ...String: MAAAAAEEGM EPRALQYEQT LMYGRYTQDL GAFAKEEAAR IRLGGPEPWK GPPSSRAAPE LLEYGRSRCA RCRVCSVRCH KFLVSRVGE DWIFLVLLGL LMALVSWVMD YAIAACLQAQ QWMSRGLNTS ILLQYLAWVT YPVVLITFSA GFTQILAPQA V GSGIPEMK TILRGVVLKE YLTLKTFIAK VIGLTCALGS GMPLGKEGPF VHIASMCAAL LSKFLSLFGG IYENESRNTE ML AAACAVG VGCCFAAPIG GVLFSIEVTS TFFAVRNYWR GFFAATFSAF IFRVLAVWNR DEETITALFK TRFRLDFPFD LQE LPAFAV IGIASGFGGA LFVYLNRKIV QVMRKQKTIN RFLMRKRLLF PALVTLLIST LTFPPGFGQF MAGQLSQKET LVTL FDNRT WVRQGLVEEL EPPSTSQAWN PPRANVFLTL VIFILMKFWM SALATTIPVP CGAFMPVFVI GAAFGRLVGE SMAAW FPDG IHTDSSTYRI VPGGYAVVGA AALAGAVTHT VSTAVIVFEL TGQIAHILPV MIAVILANAV AQSLQPSLYD SIIRIK KLP YLPELGWGRH QQYRVRVEDI MVRDVPHVAL SCTFRDLRLA LHRTKGRMLA LVESPESMIL LGSIERSQVV ALLGAQL SP ARRRQHMQER RATQTSPLSD QEGPPTPEAS VCFQVNTEDS AFPAARGETH KPLKPALKRG PSVTRNLGES PTGSAESA G IALRSLFCGS PPPEAASEKL ESCEKRKLKR VRISLASDAD LEGEMSPEEI LEWEEQQLDE PVNFSDCKID PAPFQLVER TSLHKTHTIF SLLGVDHAYV TSIGRLIGIV TLKELRKAIE GSVTAQGVKV RPPLASFRDS ATSSSDTETT EVHALWGPHS RHGLPREGS PSDSDDKCQ UniProtKB: Chloride channel protein 2 |
-Macromolecule #2: 2-[[2,6-bis(chloranyl)-3-phenylmethoxy-phenyl]amino]pyridine-3-ca...
| Macromolecule | Name: 2-[[2,6-bis(chloranyl)-3-phenylmethoxy-phenyl]amino]pyridine-3-carboxylic acid type: ligand / ID: 2 / Number of copies: 2 / Formula: GH6 |
|---|---|
| Molecular weight | Theoretical: 389.232 Da |
| Chemical component information | 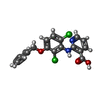 ChemComp-GH6: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8gqu: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)