[English] 日本語
 Yorodumi
Yorodumi- EMDB-34054: Cryo-EM structure of compact CA16 empty particle in complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of compact CA16 empty particle in complex with a neutralizing antibody 8C4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | coxsackievirus A16 / antibody 8C4 / cryo-EM / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / virion component / ribonucleoside triphosphate phosphatase activity / host cell ...symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / virion component / ribonucleoside triphosphate phosphatase activity / host cell / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / DNA replication / RNA helicase activity / endocytosis involved in viral entry into host cell / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / symbiont entry into host cell / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding Similarity search - Function | |||||||||
| Biological species |   Coxsackievirus A16 Coxsackievirus A16 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.91 Å | |||||||||
 Authors Authors | Cong Y / Liu CX | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Molecular mechanism of antibody neutralization of coxsackievirus A16. Authors: Chao Zhang / Caixuan Liu / Jinping Shi / Yalei Wang / Cong Xu / Xiaohua Ye / Qingwei Liu / Xue Li / Weihua Qiao / Yannan Yin / Yao Cong / Zhong Huang /  Abstract: Coxsackievirus A16 (CVA16) causes hand, foot and mouth disease in infants and young children. However, no vaccine or anti-viral agent is currently available for CVA16. Here, the functions and working ...Coxsackievirus A16 (CVA16) causes hand, foot and mouth disease in infants and young children. However, no vaccine or anti-viral agent is currently available for CVA16. Here, the functions and working mechanisms of two CVA16-specific neutralizing monoclonal antibodies (MAbs), 9B5 and 8C4, are comprehensively investigated. Both 9B5 and 8C4 display potent neutralization in vitro and prophylactic and therapeutic efficacy in a mouse model of CVA16 infection. Mechanistically, 9B5 exerts neutralization primarily through inhibiting CVA16 attachment to cell surface via blockade of CVA16 binding to its attachment receptor, heparan sulfate, whereas 8C4 functions mainly at the post-attachment stage of CVA16 entry by interfering with the interaction between CVA16 and its uncoating receptor SCARB2. Cryo-EM studies show that 9B5 and 8C4 target distinct epitopes located at the 5-fold and 3-fold protrusions of CVA16 capsids, respectively, and exhibit differential binding preference to three forms of naturally occurring CVA16 particles. Moreover, 9B5 and 8C4 are compatible in formulating an antibody cocktail which displays the ability to prevent virus escape seen with individual MAbs. Together, our work elucidates the functional and structural basis of CVA16 antibody-mediated neutralization and protection, providing important information for design and development of effective CVA16 vaccines and antibody therapies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34054.map.gz emd_34054.map.gz | 478.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34054-v30.xml emd-34054-v30.xml emd-34054.xml emd-34054.xml | 14.7 KB 14.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34054.png emd_34054.png | 49.4 KB | ||
| Filedesc metadata |  emd-34054.cif.gz emd-34054.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34054 http://ftp.pdbj.org/pub/emdb/structures/EMD-34054 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34054 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34054 | HTTPS FTP |
-Related structure data
| Related structure data |  7yrfMC  7y7mC  7ymsC  7yrhC  7yv2C  7yv7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34054.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34054.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
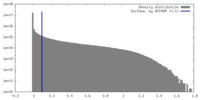
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM structure of compact CA16 empty particle in complex with ...
| Entire | Name: Cryo-EM structure of compact CA16 empty particle in complex with a neutralizing antibody 8C4 |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of compact CA16 empty particle in complex with ...
| Supramolecule | Name: Cryo-EM structure of compact CA16 empty particle in complex with a neutralizing antibody 8C4 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
-Macromolecule #1: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
| Molecular weight | Theoretical: 25.623205 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HSTQETAIGN FFSRAGLVSI ITMPTMGTQN TDGYANWDID LMGYAQLRRK CELFTYMRFD AEFTFVVAKP NGELVPQLLQ YMYVPPGAP KPTSRDSFAW QTATNPSVFV KMTDPPAQVS VPFMSPASAY QWFYDGYPTF GEHLQANDLD YGQCPNNMMG T FSIRTVGT ...String: HSTQETAIGN FFSRAGLVSI ITMPTMGTQN TDGYANWDID LMGYAQLRRK CELFTYMRFD AEFTFVVAKP NGELVPQLLQ YMYVPPGAP KPTSRDSFAW QTATNPSVFV KMTDPPAQVS VPFMSPASAY QWFYDGYPTF GEHLQANDLD YGQCPNNMMG T FSIRTVGT KKSPHSITLR VYMRIKHVRA WIPRPLRNQP YLFKTNPNYK GNDIKCTSTS RDKITTL UniProtKB: Genome polyprotein |
-Macromolecule #2: Light chain of chain
| Macromolecule | Name: Light chain of chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
| Molecular weight | Theoretical: 23.619988 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIQMTQSSSY LSVSLGGRVT ITCKASDHIN NWLAWYQQKP GNAPRLLISG ATSLETGVPS RFSGSGSGKD YTLSITSLQT EDVATYYCQ QYWNSPYTFG GGTKLEIKRA DAAPTVSIFP PSSEQLTSGG ASVVCFLNNF YPKDINVKWK IDGSERQNGV L NSWTDQDS ...String: DIQMTQSSSY LSVSLGGRVT ITCKASDHIN NWLAWYQQKP GNAPRLLISG ATSLETGVPS RFSGSGSGKD YTLSITSLQT EDVATYYCQ QYWNSPYTFG GGTKLEIKRA DAAPTVSIFP PSSEQLTSGG ASVVCFLNNF YPKDINVKWK IDGSERQNGV L NSWTDQDS KDSTYSMSST LTLTKDEYER HNSYTCEATH KTSTSPIVKS FNRNEC |
-Macromolecule #3: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: picornain 2A |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
| Molecular weight | Theoretical: 33.77284 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ENSNSASEGS TINYTTINYY KDAYAASAGR QDMSQDPKRF TDPVMDVIHE MAPPLKSPSA EACGYSDRVA QLTIGNSTIT TQEAANIVI AYGEWPEYCP DTDATAVDKP TRPDVSVNRF FTLDTKSWAK DSKGWYWKFP DVLTEVGVFG QNAQFHYLYR S GFCVHVQC ...String: ENSNSASEGS TINYTTINYY KDAYAASAGR QDMSQDPKRF TDPVMDVIHE MAPPLKSPSA EACGYSDRVA QLTIGNSTIT TQEAANIVI AYGEWPEYCP DTDATAVDKP TRPDVSVNRF FTLDTKSWAK DSKGWYWKFP DVLTEVGVFG QNAQFHYLYR S GFCVHVQC NASKFHQGAL LVAVLPEYVL GTIAGGTGNE NSHPPYATTQ PGQVGAVLTH PYVLDAGIPL SQLTVCPHQW IN LRTNNCA TIIVPYMNTV PFDSALNHCN FGLLVIPVVP LDFNAGATSE IPITVTIAPM CAEFAGLRQA VKQ UniProtKB: Genome polyprotein |
-Macromolecule #4: The heavy chain of the antibody 8C4
| Macromolecule | Name: The heavy chain of the antibody 8C4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
| Molecular weight | Theoretical: 22.743381 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLQQSGPE LVKPGASVKI SCKASGYAFS TSWMNWVIQR PGQGLEWIGR IYPGDGDTNY NGKFKGKATL TADKSSSTAY MQLSSLTSV DSAVYFCARR DYGYFDYWGQ GTTLTVSSAK TTPPSVYPLA PGCGDTTGSS VTLGCLVKGY FPESVTVTWN S GSLSSSVH ...String: QVQLQQSGPE LVKPGASVKI SCKASGYAFS TSWMNWVIQR PGQGLEWIGR IYPGDGDTNY NGKFKGKATL TADKSSSTAY MQLSSLTSV DSAVYFCARR DYGYFDYWGQ GTTLTVSSAK TTPPSVYPLA PGCGDTTGSS VTLGCLVKGY FPESVTVTWN S GSLSSSVH TFPALLQSGL YTMSSSVTVP SSTWPSQTVT CSVAHPASST TVDKKL |
-Macromolecule #5: Capsid protein VP3
| Macromolecule | Name: Capsid protein VP3 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO / EC number: picornain 2A |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A16 Coxsackievirus A16 |
| Molecular weight | Theoretical: 26.646318 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GIPTELKPGT NQFLTTDDGV SAPILPGFHP TPPIHIPGEV RNLLEICRVE TILEVNNLKT NETTPMQRLC FPVSVQSKTG ELCAAFRAD PGRDGPWQST ILGQLCRYYT QWSGSLEVTF MFAGSFMATG KMLIAYTPPG GSVPADRITA MLGTHVIWDF G LQSSVTLV ...String: GIPTELKPGT NQFLTTDDGV SAPILPGFHP TPPIHIPGEV RNLLEICRVE TILEVNNLKT NETTPMQRLC FPVSVQSKTG ELCAAFRAD PGRDGPWQST ILGQLCRYYT QWSGSLEVTF MFAGSFMATG KMLIAYTPPG GSVPADRITA MLGTHVIWDF G LQSSVTLV VPWISNTHYR AHARAGYFDY YTTGIITIWY QTNYVVPIGA PTTAYIVALA AAQDNFTMKL CKDTEDIEQT AN IQ UniProtKB: Genome polyprotein |
-Macromolecule #6: SPHINGOSINE
| Macromolecule | Name: SPHINGOSINE / type: ligand / ID: 6 / Number of copies: 1 / Formula: SPH |
|---|---|
| Molecular weight | Theoretical: 299.492 Da |
| Chemical component information |  ChemComp-SPH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.91 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 20196 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)