[English] 日本語
 Yorodumi
Yorodumi- EMDB-33522: The pre-fusion structure of Thogotovirus dhori envelope glycoprotein -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The pre-fusion structure of Thogotovirus dhori envelope glycoprotein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Viral surface protein / STRUCTURAL PROTEIN | |||||||||
| Function / homology | Baculovirus Gp64, envelope glycoprotein / Baculovirus gp64 envelope glycoprotein / symbiont-mediated perturbation of host process / viral envelope / virion membrane / membrane / Envelope glycoprotein Function and homology information Function and homology information | |||||||||
| Biological species |  Dhori thogotovirus Dhori thogotovirus | |||||||||
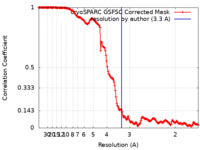
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Shan YY / Zhang MF / Pei DQ | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Pre- and post-fusion class III glycoprotein structures reveal viral fusion mechanism Authors: Shan YY / Zhang MF / Pei DQ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33522.map.gz emd_33522.map.gz | 398.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33522-v30.xml emd-33522-v30.xml emd-33522.xml emd-33522.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33522_fsc.xml emd_33522_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_33522.png emd_33522.png | 33.8 KB | ||
| Filedesc metadata |  emd-33522.cif.gz emd-33522.cif.gz | 5.1 KB | ||
| Others |  emd_33522_half_map_1.map.gz emd_33522_half_map_1.map.gz emd_33522_half_map_2.map.gz emd_33522_half_map_2.map.gz | 391.9 MB 391.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33522 http://ftp.pdbj.org/pub/emdb/structures/EMD-33522 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33522 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33522 | HTTPS FTP |
-Related structure data
| Related structure data |  7xymMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33522.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33522.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.849 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33522_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
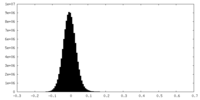
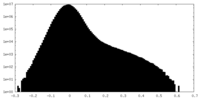
| Density Histograms |
-Half map: #2
| File | emd_33522_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
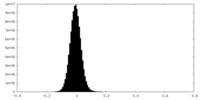
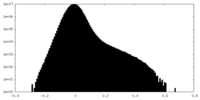
| Density Histograms |
- Sample components
Sample components
-Entire : Thogotovirus dhori envelope glycoprotein
| Entire | Name: Thogotovirus dhori envelope glycoprotein |
|---|---|
| Components |
|
-Supramolecule #1: Thogotovirus dhori envelope glycoprotein
| Supramolecule | Name: Thogotovirus dhori envelope glycoprotein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Dhori thogotovirus Dhori thogotovirus |
-Macromolecule #1: Envelope glycoprotein
| Macromolecule | Name: Envelope glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Dhori thogotovirus / Strain: Indian/1313/61 Dhori thogotovirus / Strain: Indian/1313/61 |
| Molecular weight | Theoretical: 58.674496 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDSTIRLVAT IFLISLTQQI EVCNKAQQQG PYTLVDYQEK PLNISRIQIK VVKTSVATKG LNFHIGYRAV WRSYCYNGGS LDKNTGCYN DLIPKSPTES ELRTWSKSQK CCTGPDAVDA WGSDARICWA EWKMELCHTA KELKKYSNNN HFAYHTCNLS W RCGLKSTH ...String: MDSTIRLVAT IFLISLTQQI EVCNKAQQQG PYTLVDYQEK PLNISRIQIK VVKTSVATKG LNFHIGYRAV WRSYCYNGGS LDKNTGCYN DLIPKSPTES ELRTWSKSQK CCTGPDAVDA WGSDARICWA EWKMELCHTA KELKKYSNNN HFAYHTCNLS W RCGLKSTH IEVRLQASGG LVSMVAVMPN GTLIPIEGTR PTYWTEDSFA YLYDPAGTEK KTESTFLWCF KEHIRPTTEL SG AVYDTHY LGGTYDKNPQ FNYYCRDNGY YFELPANRLV CLPTSCYKRE GAIVNTMHPD TWKVSEKLHS ASQFDVNNVV HSL VYETEG LRLALSQLDH RFATLSRLFN RLTQSLAKID DRLLGTLLGQ DVSSKFISPT KFMLSPCLST PEGDSNCHNH SIYR DGRWV HNSDPTQCFS LSKSQPVDLY SFKELWLPQL LDVNVEGVVA DEEGWSFVAQ SKQALIDTMT YTKNGGKGTS LEDVL GYPS GWINGKLQGL LLNGAISWVV VIGVVLVGVC LMRRVF UniProtKB: Envelope glycoprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)