+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | High resolution cry-EM structure of the human 80S ribosome from SNORD127+/+ Kasumi-1 cells | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報positive regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis / negative regulation of endoplasmic reticulum unfolded protein response / oxidized pyrimidine DNA binding / eukaryotic 80S initiation complex / response to TNF agonist / positive regulation of base-excision repair / negative regulation of protein neddylation / protein tyrosine kinase inhibitor activity / positive regulation of respiratory burst involved in inflammatory response / translation at presynapse ...positive regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis / negative regulation of endoplasmic reticulum unfolded protein response / oxidized pyrimidine DNA binding / eukaryotic 80S initiation complex / response to TNF agonist / positive regulation of base-excision repair / negative regulation of protein neddylation / protein tyrosine kinase inhibitor activity / positive regulation of respiratory burst involved in inflammatory response / translation at presynapse / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / positive regulation of gastrulation / axial mesoderm development / negative regulation of formation of translation preinitiation complex / nucleolus organization / ribosomal protein import into nucleus / IRE1-RACK1-PP2A complex / : / positive regulation of endodeoxyribonuclease activity / positive regulation of Golgi to plasma membrane protein transport / 90S preribosome assembly / TNFR1-mediated ceramide production / negative regulation of RNA splicing / negative regulation of DNA repair / TORC2 complex binding / GAIT complex / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / oxidized purine DNA binding / supercoiled DNA binding / neural crest cell differentiation / NF-kappaB complex / middle ear morphogenesis / ubiquitin-like protein conjugating enzyme binding / regulation of establishment of cell polarity / negative regulation of phagocytosis / positive regulation of ubiquitin-protein transferase activity / Formation of the ternary complex, and subsequently, the 43S complex / rRNA modification in the nucleus and cytosol / erythrocyte homeostasis / cytoplasmic side of rough endoplasmic reticulum membrane / A band / laminin receptor activity / regulation of G1 to G0 transition / exit from mitosis / alpha-beta T cell differentiation / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of translation involved in cellular response to UV / protein kinase A binding / protein-DNA complex disassembly / positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / Translation initiation complex formation / Ribosomal scanning and start codon recognition / negative regulation of ubiquitin protein ligase activity / optic nerve development / ion channel inhibitor activity / pigmentation / mammalian oogenesis stage / positive regulation of mitochondrial depolarization / retinal ganglion cell axon guidance / G1 to G0 transition / homeostatic process / activation-induced cell death of T cells / lung morphogenesis / negative regulation of Wnt signaling pathway / fibroblast growth factor binding / positive regulation of T cell receptor signaling pathway / positive regulation of activated T cell proliferation / iron-sulfur cluster binding / regulation of cell division / Protein hydroxylation / negative regulation of peptidyl-serine phosphorylation / BH3 domain binding / mTORC1-mediated signalling / macrophage chemotaxis / SARS-CoV-1 modulates host translation machinery / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / Peptide chain elongation / monocyte chemotaxis / Selenocysteine synthesis / cysteine-type endopeptidase activator activity involved in apoptotic process / positive regulation of signal transduction by p53 class mediator / ubiquitin ligase inhibitor activity / Formation of a pool of free 40S subunits / phagocytic cup / Eukaryotic Translation Termination / blastocyst development / Response of EIF2AK4 (GCN2) to amino acid deficiency / SRP-dependent cotranslational protein targeting to membrane / negative regulation of respiratory burst involved in inflammatory response / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Viral mRNA Translation / protein localization to nucleus / negative regulation of proteasomal ubiquitin-dependent protein catabolic process / L13a-mediated translational silencing of Ceruloplasmin expression / GTP hydrolysis and joining of the 60S ribosomal subunit / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / TOR signaling / T cell proliferation involved in immune response / Major pathway of rRNA processing in the nucleolus and cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  human (ヒト) human (ヒト) | |||||||||
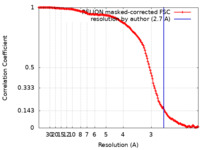
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.7 Å | |||||||||
 データ登録者 データ登録者 | Cheng J / Beckmann R | |||||||||
| 資金援助 | 1件
| |||||||||
 引用 引用 |  ジャーナル: Cancer Discov / 年: 2023 ジャーナル: Cancer Discov / 年: 2023タイトル: A Dynamic rRNA Ribomethylome Drives Stemness in Acute Myeloid Leukemia. 著者: Fengbiao Zhou / Nesrine Aroua / Yi Liu / Christian Rohde / Jingdong Cheng / Anna-Katharina Wirth / Daria Fijalkowska / Stefanie Göllner / Michelle Lotze / Haiyang Yun / Xiaobing Yu / ...著者: Fengbiao Zhou / Nesrine Aroua / Yi Liu / Christian Rohde / Jingdong Cheng / Anna-Katharina Wirth / Daria Fijalkowska / Stefanie Göllner / Michelle Lotze / Haiyang Yun / Xiaobing Yu / Caroline Pabst / Tim Sauer / Thomas Oellerich / Hubert Serve / Christoph Röllig / Martin Bornhäuser / Christian Thiede / Claudia Baldus / Michaela Frye / Simon Raffel / Jeroen Krijgsveld / Irmela Jeremias / Roland Beckmann / Andreas Trumpp / Carsten Müller-Tidow /  要旨: The development and regulation of malignant self-renewal remain unresolved issues. Here, we provide biochemical, genetic, and functional evidence that dynamics in ribosomal RNA (rRNA) 2'-O- ...The development and regulation of malignant self-renewal remain unresolved issues. Here, we provide biochemical, genetic, and functional evidence that dynamics in ribosomal RNA (rRNA) 2'-O-methylation regulate leukemia stem cell (LSC) activity in vivo. A comprehensive analysis of the rRNA 2'-O-methylation landscape of 94 patients with acute myeloid leukemia (AML) revealed dynamic 2'-O-methylation specifically at exterior sites of ribosomes. The rRNA 2'-O-methylation pattern is closely associated with AML development stage and LSC gene expression signature. Forced expression of the 2'-O-methyltransferase fibrillarin (FBL) induced an AML stem cell phenotype and enabled engraftment of non-LSC leukemia cells in NSG mice. Enhanced 2'-O-methylation redirected the ribosome translation program toward amino acid transporter mRNAs enriched in optimal codons and subsequently increased intracellular amino acid levels. Methylation at the single site 18S-guanosine 1447 was instrumental for LSC activity. Collectively, our work demonstrates that dynamic 2'-O-methylation at specific sites on rRNAs shifts translational preferences and controls AML LSC self-renewal. SIGNIFICANCE: We establish the complete rRNA 2'-O-methylation landscape in human AML. Plasticity of rRNA 2'-O-methylation shifts protein translation toward an LSC phenotype. This dynamic process ...SIGNIFICANCE: We establish the complete rRNA 2'-O-methylation landscape in human AML. Plasticity of rRNA 2'-O-methylation shifts protein translation toward an LSC phenotype. This dynamic process constitutes a novel concept of how cancers reprogram cell fate and function. This article is highlighted in the In This Issue feature, p. 247. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_33329.map.gz emd_33329.map.gz | 157.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-33329-v30.xml emd-33329-v30.xml emd-33329.xml emd-33329.xml | 95.5 KB 95.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_33329_fsc.xml emd_33329_fsc.xml | 14.8 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_33329.png emd_33329.png | 236.9 KB | ||
| その他 |  emd_33329_additional_1.map.gz emd_33329_additional_1.map.gz emd_33329_half_map_1.map.gz emd_33329_half_map_1.map.gz emd_33329_half_map_2.map.gz emd_33329_half_map_2.map.gz | 15.8 MB 225.2 MB 225.2 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33329 http://ftp.pdbj.org/pub/emdb/structures/EMD-33329 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33329 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33329 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_33329_validation.pdf.gz emd_33329_validation.pdf.gz | 1005.8 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_33329_full_validation.pdf.gz emd_33329_full_validation.pdf.gz | 1005.4 KB | 表示 | |
| XML形式データ |  emd_33329_validation.xml.gz emd_33329_validation.xml.gz | 22.7 KB | 表示 | |
| CIF形式データ |  emd_33329_validation.cif.gz emd_33329_validation.cif.gz | 30 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33329 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33329 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33329 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33329 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7xnxMC  7xnyC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_33329.map.gz / 形式: CCP4 / 大きさ: 282.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_33329.map.gz / 形式: CCP4 / 大きさ: 282.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.059 Å | ||||||||||||||||||||
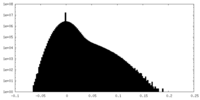
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: #1
| ファイル | emd_33329_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
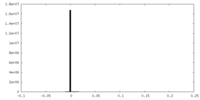
| 投影像・断面図 |
| ||||||||||||
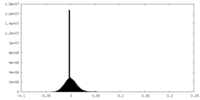
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_33329_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
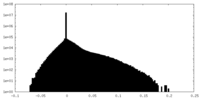
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_33329_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
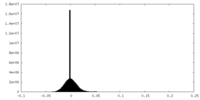
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : ribosome
+超分子 #1: ribosome
+分子 #1: 28S rRNA
+分子 #2: 5S rRNA
+分子 #3: 5.8S rRNA
+分子 #45: 18S rRNA
+分子 #79: tRNA
+分子 #4: 60S ribosomal protein L8
+分子 #5: 60S ribosomal protein L3
+分子 #6: 60S ribosomal protein L4
+分子 #7: 60S ribosomal protein L5
+分子 #8: 60S ribosomal protein L6
+分子 #9: 60S ribosomal protein L7
+分子 #10: 60S ribosomal protein L7a
+分子 #11: 60S ribosomal protein L9
+分子 #12: Ribosomal protein L10 isoform A
+分子 #13: 60S ribosomal protein L11
+分子 #14: 60S ribosomal protein L13
+分子 #15: 60S ribosomal protein L14
+分子 #16: 60S ribosomal protein L15
+分子 #17: 60S ribosomal protein L13a
+分子 #18: 60S ribosomal protein L17
+分子 #19: 60S ribosomal protein L18
+分子 #20: 60S ribosomal protein L19
+分子 #21: 60S ribosomal protein L18a
+分子 #22: 60S ribosomal protein L21
+分子 #23: 60S ribosomal protein L22
+分子 #24: 60S ribosomal protein L23
+分子 #25: 60S ribosomal protein L24
+分子 #26: 60S ribosomal protein L23a
+分子 #27: 60S ribosomal protein L26
+分子 #28: 60S ribosomal protein L27
+分子 #29: 60S ribosomal protein L29
+分子 #30: 60S ribosomal protein L30
+分子 #31: 60S ribosomal protein L31
+分子 #32: 60S ribosomal protein L32
+分子 #33: 60S ribosomal protein L35a
+分子 #34: 60S ribosomal protein L34
+分子 #35: 60S ribosomal protein L35
+分子 #36: 60S ribosomal protein L36
+分子 #37: 60S ribosomal protein L37
+分子 #38: 60S ribosomal protein L38
+分子 #39: 60S ribosomal protein L39
+分子 #40: Ubiquitin-60S ribosomal protein L40
+分子 #41: 60S ribosomal protein L41
+分子 #42: 60S ribosomal protein L36a
+分子 #43: 60S ribosomal protein L37a
+分子 #44: 60S ribosomal protein L28
+分子 #46: 40S ribosomal protein SA
+分子 #47: 40S ribosomal protein S3a
+分子 #48: 40S ribosomal protein S3
+分子 #49: 40S ribosomal protein S4, X isoform
+分子 #50: 40S ribosomal protein S5
+分子 #51: 40S ribosomal protein S7
+分子 #52: 40S ribosomal protein S8
+分子 #53: 40S ribosomal protein S10
+分子 #54: 40S ribosomal protein S11
+分子 #55: 40S ribosomal protein S15
+分子 #56: 40S ribosomal protein S16
+分子 #57: 40S ribosomal protein S17
+分子 #58: 40S ribosomal protein S18
+分子 #59: 40S ribosomal protein S19
+分子 #60: 40S ribosomal protein S20
+分子 #61: 40S ribosomal protein S21
+分子 #62: 40S ribosomal protein S23
+分子 #63: 40S ribosomal protein S26
+分子 #64: 40S ribosomal protein S28
+分子 #65: 40S ribosomal protein S29
+分子 #66: Receptor of activated protein C kinase 1
+分子 #67: 40S ribosomal protein S2
+分子 #68: 40S ribosomal protein S6
+分子 #69: 40S ribosomal protein S9
+分子 #70: 40S ribosomal protein S12
+分子 #71: 40S ribosomal protein S13
+分子 #72: 40S ribosomal protein S14
+分子 #73: 40S ribosomal protein S15a
+分子 #74: 40S ribosomal protein S24
+分子 #75: 40S ribosomal protein S25
+分子 #76: 40S ribosomal protein S27
+分子 #77: 40S ribosomal protein S30
+分子 #78: Ubiquitin-40S ribosomal protein S27a
+分子 #80: MAGNESIUM ION
+分子 #81: POTASSIUM ION
+分子 #82: ZINC ION
+分子 #83: water
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / 平均電子線量: 44.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源: TUNGSTEN HAIRPIN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 0.8 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー





































 Z
Z Y
Y X
X