[English] 日本語
 Yorodumi
Yorodumi- EMDB-33321: Cryo-EM structure of the bacteriocin-receptor-immunity ternary co... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the bacteriocin-receptor-immunity ternary complex from Lactobacillus sakei | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationbacteriocin immunity / cytolysis / phosphoenolpyruvate-dependent sugar phosphotransferase system / defense response to bacterium / extracellular region / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) | |||||||||
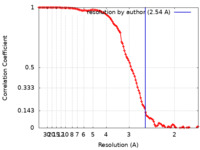
| Method | single particle reconstruction / cryo EM / Resolution: 2.54 Å | |||||||||
 Authors Authors | Wang JW | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Appl Environ Microbiol / Year: 2022 Journal: Appl Environ Microbiol / Year: 2022Title: Structural Basis of the Immunity Mechanisms of Pediocin-like Bacteriocins. Authors: Liyan Zhu / Jianwei Zeng / Jiawei Wang /  Abstract: Pediocin-like bacteriocins, also designated class IIa bacteriocins, are ribosomally synthesized antimicrobial peptides targeting species closely related to the producers. They act on the cytoplasmic ...Pediocin-like bacteriocins, also designated class IIa bacteriocins, are ribosomally synthesized antimicrobial peptides targeting species closely related to the producers. They act on the cytoplasmic membrane of Gram-positive cells by dissipating the transmembrane electrical potential through pore formation with the mannose phosphotransferase system (man-PTS) as the target/receptor. Bacteriocin-producing strains also synthesize a cognate immunity protein that protects them against their own bacteriocins. Herein, we report the cryo-electron microscopy structure of the bacteriocin-receptor-immunity ternary complex from Lactobacillus sakei. The complex structure reveals that pediocin-like bacteriocins bind to the same position on the Core domain of man-PTS, while the C-terminal helical tails of bacteriocins delimit the opening range of the Core domain away from the Vmotif domain to facilitate transmembrane pore formation. Upon attack of bacteriocins from the extracellular side, man-PTS exposes its cytosolic side for recognition of the N-terminal four-helix bundle of the immunity protein. The C-terminal loop of the immunity protein then inserts into the pore and blocks leakage induced by bacteriocins. Elucidation of the toxicity and immunity mechanisms of pediocin-like bacteriocins could support the design of novel bacteriocins against antibiotic-resistant pathogenic bacteria. Pediocin-like bacteriocins, ribosomally synthesized antimicrobial peptides, are generally co-expressed with cognate immunity proteins to protect the bacteriocin-producing strain from its own bacteriocin. Bacteriocins are considered potential alternatives to conventional antibiotics in the context of the bacterial resistance crisis, but the immunity mechanism is unclear. This study uncovered the mechanisms of action and immunity of class IIa bacteriocins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33321.map.gz emd_33321.map.gz | 117.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33321-v30.xml emd-33321-v30.xml emd-33321.xml emd-33321.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33321_fsc.xml emd_33321_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_33321.png emd_33321.png | 35 KB | ||
| Others |  emd_33321_half_map_1.map.gz emd_33321_half_map_1.map.gz emd_33321_half_map_2.map.gz emd_33321_half_map_2.map.gz | 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33321 http://ftp.pdbj.org/pub/emdb/structures/EMD-33321 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33321 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33321 | HTTPS FTP |
-Validation report
| Summary document |  emd_33321_validation.pdf.gz emd_33321_validation.pdf.gz | 780.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33321_full_validation.pdf.gz emd_33321_full_validation.pdf.gz | 779.9 KB | Display | |
| Data in XML |  emd_33321_validation.xml.gz emd_33321_validation.xml.gz | 19.2 KB | Display | |
| Data in CIF |  emd_33321_validation.cif.gz emd_33321_validation.cif.gz | 24.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33321 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33321 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33321 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33321 | HTTPS FTP |
-Related structure data
| Related structure data |  7xnoMC  7xtgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33321.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33321.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.8433 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map 1
| File | emd_33321_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
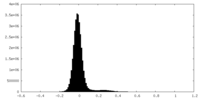
| Density Histograms |
-Half map: Half map 2
| File | emd_33321_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
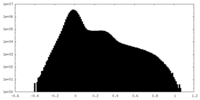
| Density Histograms |
- Sample components
Sample components
-Entire : The bacteriocin-receptor-immunity ternary complex from Lactobacil...
| Entire | Name: The bacteriocin-receptor-immunity ternary complex from Lactobacillus sakei |
|---|---|
| Components |
|
-Supramolecule #1: The bacteriocin-receptor-immunity ternary complex from Lactobacil...
| Supramolecule | Name: The bacteriocin-receptor-immunity ternary complex from Lactobacillus sakei type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) |
| Recombinant expression | Organism:  |
-Macromolecule #1: Bacteriocin sakacin-A
| Macromolecule | Name: Bacteriocin sakacin-A / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) |
| Molecular weight | Theoretical: 4.312893 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ARSYGNGVYC NNKKCWVNRG EATQSIIGGM ISGWASGLAG M |
-Macromolecule #2: Mannose permease IIC component
| Macromolecule | Name: Mannose permease IIC component / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) |
| Molecular weight | Theoretical: 27.467719 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDLNFIQVIL VIFVAFLAGV EGILDQFHFH QPVIACTLIG LVTGNLLPCL ILGGTLQMIA LGWANVGAAV APDAALASIA SAIILVLGG QGKAGVTSAI AIAVPLAVAG LLLTIIVRTL ATGIVHIMDA AAKEGNFRKI EMWQYIAIIM QGVRIAIPAG L ILAIGAGP ...String: MDLNFIQVIL VIFVAFLAGV EGILDQFHFH QPVIACTLIG LVTGNLLPCL ILGGTLQMIA LGWANVGAAV APDAALASIA SAIILVLGG QGKAGVTSAI AIAVPLAVAG LLLTIIVRTL ATGIVHIMDA AAKEGNFRKI EMWQYIAIIM QGVRIAIPAG L ILAIGAGP VKEMLTAMPV WLTDGLAIGG GMVVAVGYAM VINMMATKEV WPFFAIGFVL ATISQLTLIG LGAIGISLAL IY LALSKQG SGNNGGGSNT GDPLGDIIDN Y |
-Macromolecule #3: Mannose permease IID component
| Macromolecule | Name: Mannose permease IID component / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) |
| Molecular weight | Theoretical: 33.332105 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAEQLKLTKK DRISVWLRST FLQGSWNYER MQNGGWAYTL IPALKKLYKT KEDRSAALVR HMEFFNTHPY VAAPILGVTL ALEEERANG APIDDVTIQG VKVGMMGPLA GIGDPVFWFT VKPIIGALAA SLAMSGNILG PIIYFVAWNA IRMAFTWYTQ E FGYRAGSK ...String: MAEQLKLTKK DRISVWLRST FLQGSWNYER MQNGGWAYTL IPALKKLYKT KEDRSAALVR HMEFFNTHPY VAAPILGVTL ALEEERANG APIDDVTIQG VKVGMMGPLA GIGDPVFWFT VKPIIGALAA SLAMSGNILG PIIYFVAWNA IRMAFTWYTQ E FGYRAGSK ITEDLSGGIL QDITKGASIL GMFILGSLVN RWVSVKFTPT VSSVKLDKGA FIDWDKLPSG AKGIQSALQQ QA QGLSLTD HKITTLQDNL DSLIPGLAAL GLTLFCMWLL KKKVSPIVII LGLFVVGIVF HLLHLM |
-Macromolecule #4: Sakacin-A immunity factor
| Macromolecule | Name: Sakacin-A immunity factor / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Latilactobacillus sakei L45 (bacteria) Latilactobacillus sakei L45 (bacteria) |
| Molecular weight | Theoretical: 13.747889 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKHHHHHHHG AAGTSLYKKA GENLYFQGSM KADYKKINSI LTYTSTALKN PKIIKDKDLV VLLTIIQEEA KQNRIFYDYK RKFRPAVTR FTIDNNFEIP DCLVKLLSAV ETPKAWSGFS |
-Macromolecule #5: alpha-D-mannopyranose
| Macromolecule | Name: alpha-D-mannopyranose / type: ligand / ID: 5 / Number of copies: 3 / Formula: MAN |
|---|---|
| Molecular weight | Theoretical: 180.156 Da |
| Chemical component information |  ChemComp-MAN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
 Movie
Movie Controller
Controller




 Z
Z Y
Y X
X