[English] 日本語
 Yorodumi
Yorodumi- EMDB-30537: Cryo-EM map of Omono River virus (Strain: LZ) full capsid with it... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30537 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of Omono River virus (Strain: LZ) full capsid with its fiber-like protrusion anchored at the five-fold vertex. | |||||||||
 Map data Map data | To show each element of the virus, maps are shown at different counter levels. The major capsid shell: 4%u03C3; Subvolume of the protrusion: 0.65%u03C3. | |||||||||
 Sample Sample | Major capsid dimer of OmRV-LZ (with protrusion) != Omono River virus Major capsid dimer of OmRV-LZ (with protrusion)
| |||||||||
 Keywords Keywords | Totiviridae / major capsid / virus | |||||||||
| Function / homology | Double-stranded RNA binding motif / Double-stranded RNA binding motif / Double stranded RNA-binding domain (dsRBD) profile. / Double-stranded RNA-binding domain / RNA binding / Main capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Omono River virus Omono River virus | |||||||||
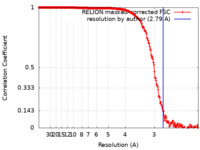
| Method | single particle reconstruction / cryo EM / Resolution: 2.79 Å | |||||||||
 Authors Authors | Shao Q / Jia X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2021 Journal: PLoS Pathog / Year: 2021Title: Cryo-EM reveals a previously unrecognized structural protein of a dsRNA virus implicated in its extracellular transmission. Authors: Qianqian Shao / Xudong Jia / Yuanzhu Gao / Zhe Liu / Huan Zhang / Qiqi Tan / Xin Zhang / Huiqiong Zhou / Yinyin Li / De Wu / Qinfen Zhang /  Abstract: Mosquito viruses cause unpredictable outbreaks of disease. Recently, several unassigned viruses isolated from mosquitoes, including the Omono River virus (OmRV), were identified as totivirus-like ...Mosquito viruses cause unpredictable outbreaks of disease. Recently, several unassigned viruses isolated from mosquitoes, including the Omono River virus (OmRV), were identified as totivirus-like viruses, with features similar to those of the Totiviridae family. Most reported members of this family infect fungi or protozoans and lack an extracellular life cycle stage. Here, we identified a new strain of OmRV and determined high-resolution structures for this virus using single-particle cryo-electron microscopy. The structures feature an unexpected protrusion at the five-fold vertex of the capsid. Disassociation of the protrusion could result in several conformational changes in the major capsid. All these structures, together with some biological results, suggest the protrusions' associations with the extracellular transmission of OmRV. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30537.map.gz emd_30537.map.gz | 771.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30537-v30.xml emd-30537-v30.xml emd-30537.xml emd-30537.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30537_fsc.xml emd_30537_fsc.xml | 21.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_30537.png emd_30537.png | 239.4 KB | ||
| Filedesc metadata |  emd-30537.cif.gz emd-30537.cif.gz | 6.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30537 http://ftp.pdbj.org/pub/emdb/structures/EMD-30537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30537 | HTTPS FTP |
-Related structure data
| Related structure data |  7d0kMC  7cz6C  7d0lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30537.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30537.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | To show each element of the virus, maps are shown at different counter levels. The major capsid shell: 4%u03C3; Subvolume of the protrusion: 0.65%u03C3. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.09 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Major capsid dimer of OmRV-LZ (with protrusion)
| Entire | Name: Major capsid dimer of OmRV-LZ (with protrusion) |
|---|---|
| Components |
|
-Supramolecule #1: Omono River virus
| Supramolecule | Name: Omono River virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 753758 / Sci species name: Omono River virus / Sci species strain: LZ / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Culex quinquefasciatus (southern house mosquito) Culex quinquefasciatus (southern house mosquito) |
| Virus shell | Shell ID: 1 / Name: Major capsid / Diameter: 450.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Omono River virus / Strain: LZ Omono River virus / Strain: LZ |
| Molecular weight | Theoretical: 96.860992 KDa |
| Sequence | String: PISADFSEVE NAPSFLSLAE NTDEVLKPYT GLEIQTIITN IVGDANPNQS RIFDQDRLRG NQYSAGGLVT QNAVSAIPFT NLIPRTIRV GNILVNSANR LQITETNVSE YYSNPIIATK LSEMISDQVK NNQFSTWRRD NTSLQGFNAF DIATINTAIL P NGLSLESM ...String: PISADFSEVE NAPSFLSLAE NTDEVLKPYT GLEIQTIITN IVGDANPNQS RIFDQDRLRG NQYSAGGLVT QNAVSAIPFT NLIPRTIRV GNILVNSANR LQITETNVSE YYSNPIIATK LSEMISDQVK NNQFSTWRRD NTSLQGFNAF DIATINTAIL P NGLSLESM LLKLSLLHSI KAMNVDAASI NRSQYQVIDH NTVPTIGAPA VVGVNNSPVF GEDCGGNNPV YPFGGGTGAI AF HVTLQTV PDERKSYAIF VPPAILQATS DANEALALFA LSMSEWPHAL YTVTKQTTDL AGANAGQQVF IPTQSTIHIG GRR VLDLII PRREIAPNPT TLVAANAMCM VRPQAGPDAT AGAIPLAAGQ LFNMNFIGAP AFEEWPMTSY LYSWAGRFDI TTIR QYMGR LATMVGVKDA YWAAHELNVA LSQVAPKMTT AAGGWAAQAA NSAQQSDVCY SSLLTVTRSA ANFPLANQPA ADMRV YDTD PATWNKVALG LATAANLVPE QSMDVPFVVG DARASFWERL QAIPMCIAWT MYYHSRGITT LAWDNAYTDN TNKWLQ KMV RNTFSTTQSV GTIIPARYGK IVCNLYKNMF HRAPAYVATS VGGKELHITH FERWLPGGTY ANVYSGAGAV VNCFSPV LI PDIWCQYFTA KLPLFAGAFP PAQGQNSTKG FNSKQGLMIH RNQNNNLVAP YLEKFADNSS YFPVGQGPEI NDMATWNG R LWMTTGNVQY LDYSGAAIVE AVPPAGELPV GKQIPLLAGE NAPIELTNAA TTCVPRYSND GRRIFTYLTT AQSVIPVQA CNRAANLARS CWLLSNVYAE PALQALGDEV EDAFDTLTNS SFLDVAKSVA ESAGEVPATK ALTDLQAVDV SSLPSTSDPS NVLSQPAPL MSPPTSSS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 75000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7d0k: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)