+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Yeast ATP Synthase map in presence of MgATP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | F-type / ATP Synthase / yeast / mitochondrial / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information: / : / : / : / : / cristae formation / Mitochondrial protein degradation / mitochondrial proton-transporting ATP synthase complex assembly / photosynthetic electron transport in photosystem I / photosynthetic electron transport in photosystem II ...: / : / : / : / : / cristae formation / Mitochondrial protein degradation / mitochondrial proton-transporting ATP synthase complex assembly / photosynthetic electron transport in photosystem I / photosynthetic electron transport in photosystem II / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / proton motive force-driven mitochondrial ATP synthesis / proton-transporting ATPase activity, rotational mechanism / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / proton transmembrane transport / ADP binding / mitochondrial membrane / protein-containing complex assembly / mitochondrial inner membrane / hydrolase activity / lipid binding / ATP hydrolysis activity / mitochondrion / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Sharma S / Patel H / Luo M / Mueller DM / Liao M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Conformational ensemble of yeast ATP synthase at low pH reveals unique intermediates and plasticity in F-F coupling. Authors: Stuti Sharma / Min Luo / Hiral Patel / David M Mueller / Maofu Liao /    Abstract: Mitochondrial adenosine triphosphate (ATP) synthase uses the proton gradient across the inner mitochondrial membrane to synthesize ATP. Structural and single molecule studies conducted mostly at ...Mitochondrial adenosine triphosphate (ATP) synthase uses the proton gradient across the inner mitochondrial membrane to synthesize ATP. Structural and single molecule studies conducted mostly at neutral or basic pH have provided details of the reaction mechanism of ATP synthesis. However, pH of the mitochondrial matrix is slightly acidic during hypoxia and pH-dependent conformational changes in the ATP synthase have been reported. Here we use single-particle cryo-EM to analyze the conformational ensemble of the yeast (Saccharomyces cerevisiae) ATP synthase at pH 6. Of the four conformations resolved in this study, three are reaction intermediates. In addition to canonical catalytic dwell and binding dwell structures, we identify two unique conformations with nearly identical positions of the central rotor but different catalytic site conformations. These structures provide new insights into the catalytic mechanism of the ATP synthase and highlight elastic coupling between the catalytic and proton translocating domains. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29270.map.gz emd_29270.map.gz | 49.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29270-v30.xml emd-29270-v30.xml emd-29270.xml emd-29270.xml | 31.4 KB 31.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29270.png emd_29270.png | 40.2 KB | ||
| Filedesc metadata |  emd-29270.cif.gz emd-29270.cif.gz | 8 KB | ||
| Others |  emd_29270_half_map_1.map.gz emd_29270_half_map_1.map.gz emd_29270_half_map_2.map.gz emd_29270_half_map_2.map.gz | 49.6 MB 49.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29270 http://ftp.pdbj.org/pub/emdb/structures/EMD-29270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29270 | HTTPS FTP |
-Related structure data
| Related structure data |  8fl8MC  8f29C  8f39C  8fkjC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29270.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29270.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
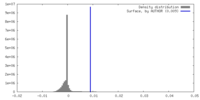
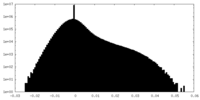
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_29270_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
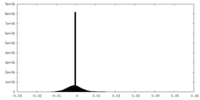
| Density Histograms |
-Half map: #2
| File | emd_29270_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
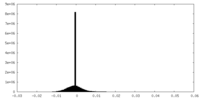
| Density Histograms |
- Sample components
Sample components
+Entire : ATP Synthase
+Supramolecule #1: ATP Synthase
+Macromolecule #1: ATP synthase protein 8
+Macromolecule #2: ATP18 isoform 1
+Macromolecule #3: ATP synthase subunit 5, mitochondrial
+Macromolecule #4: ATP synthase subunit alpha
+Macromolecule #5: ATP synthase subunit beta
+Macromolecule #6: ATP synthase subunit gamma, mitochondrial
+Macromolecule #7: ATP synthase subunit delta, mitochondrial
+Macromolecule #8: ATP synthase subunit epsilon, mitochondrial
+Macromolecule #9: ATP synthase subunit 9, mitochondrial
+Macromolecule #10: ATP synthase subunit a
+Macromolecule #11: ATP synthase subunit 4, mitochondrial
+Macromolecule #12: ATP synthase subunit d, mitochondrial
+Macromolecule #13: ATP synthase subunit H, mitochondrial
+Macromolecule #14: ATP synthase subunit f, mitochondrial
+Macromolecule #15: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #16: MAGNESIUM ION
+Macromolecule #17: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 51.07 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)