+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
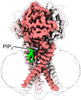
| Title | Human PAC in nanodisc at pH 4.0 with PI(4,5)P2 diC8 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | PAC / TMEM206 / ASOR / PAORAC / ion channels / chloride channel / MEMBRANE PROTEIN | ||||||||||||
| Function / homology | pH-gated chloride channel activity / TMEM206 protein / TMEM206 protein family / chloride transport / chloride channel complex / cell surface / plasma membrane / Proton-activated chloride channel Function and homology information Function and homology information | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | ||||||||||||
 Authors Authors | Ruan Z / Lu W | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2023 Journal: Elife / Year: 2023Title: Inhibition of the proton-activated chloride channel PAC by PIP. Authors: Ljubica Mihaljević / Zheng Ruan / James Osei-Owusu / Wei Lü / Zhaozhu Qiu /  Abstract: Proton-activated chloride (PAC) channel is a ubiquitously expressed pH-sensing ion channel, encoded by (). PAC regulates endosomal acidification and macropinosome shrinkage by releasing chloride ...Proton-activated chloride (PAC) channel is a ubiquitously expressed pH-sensing ion channel, encoded by (). PAC regulates endosomal acidification and macropinosome shrinkage by releasing chloride from the organelle lumens. It is also found at the cell surface, where it is activated under pathological conditions related to acidosis and contributes to acid-induced cell death. However, the pharmacology of the PAC channel is poorly understood. Here, we report that phosphatidylinositol (4,5)-bisphosphate (PIP) potently inhibits PAC channel activity. We solved the cryo-electron microscopy structure of PAC with PIP at pH 4.0 and identified its putative binding site, which, surprisingly, locates on the extracellular side of the transmembrane domain (TMD). While the overall conformation resembles the previously resolved PAC structure in the desensitized state, the TMD undergoes remodeling upon PIP-binding. Structural and electrophysiological analyses suggest that PIP inhibits the PAC channel by stabilizing the channel in a desensitized-like conformation. Our findings identify PIP as a new pharmacological tool for the PAC channel and lay the foundation for future drug discovery targeting this channel. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28964.map.gz emd_28964.map.gz | 96.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28964-v30.xml emd-28964-v30.xml emd-28964.xml emd-28964.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28964.png emd_28964.png | 126.5 KB | ||
| Masks |  emd_28964_msk_1.map emd_28964_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28964.cif.gz emd-28964.cif.gz | 6.3 KB | ||
| Others |  emd_28964_additional_1.map.gz emd_28964_additional_1.map.gz emd_28964_half_map_1.map.gz emd_28964_half_map_1.map.gz emd_28964_half_map_2.map.gz emd_28964_half_map_2.map.gz | 51.8 MB 95.4 MB 95.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28964 http://ftp.pdbj.org/pub/emdb/structures/EMD-28964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28964 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28964 | HTTPS FTP |
-Related structure data
| Related structure data |  8fblMC  8eq4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28964.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28964.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.826 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28964_msk_1.map emd_28964_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_28964_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28964_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28964_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Proton activated chloride channel at pH 4.0 with 0.5mM PIP2
| Entire | Name: Proton activated chloride channel at pH 4.0 with 0.5mM PIP2 |
|---|---|
| Components |
|
-Supramolecule #1: Proton activated chloride channel at pH 4.0 with 0.5mM PIP2
| Supramolecule | Name: Proton activated chloride channel at pH 4.0 with 0.5mM PIP2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Proton-activated chloride channel
| Macromolecule | Name: Proton-activated chloride channel / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.656955 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: FSKACLKNVF SVLLIFIYLL LMAVAVFLVY RTITDFREKL KHPVMSVSYK EVDRYDAPGI ALYPGQAQLL SCKHHYEVIP PLTSPGQPG DMNCTTQRIN YTDPFSNQTV KSALIVQGPR EVKKRELVFL QFRLNKSSED FSAIDYLLFS SFQEFLQSPN R VGFMQACE ...String: FSKACLKNVF SVLLIFIYLL LMAVAVFLVY RTITDFREKL KHPVMSVSYK EVDRYDAPGI ALYPGQAQLL SCKHHYEVIP PLTSPGQPG DMNCTTQRIN YTDPFSNQTV KSALIVQGPR EVKKRELVFL QFRLNKSSED FSAIDYLLFS SFQEFLQSPN R VGFMQACE SAYSSWKFSG GFRTWVKMSL VKTKEEDGRE AVEFRQETSV VNYIDQRPAA KKSAQLFFVV FEWKDPFIQK VQ DIVTANP WNTIALLCGA FLALFKAAEF AKLSIKWMIK IRKRYL UniProtKB: Proton-activated chloride channel |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 12 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(o...
| Macromolecule | Name: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate type: ligand / ID: 3 / Number of copies: 3 / Formula: PIO |
|---|---|
| Molecular weight | Theoretical: 746.566 Da |
| Chemical component information |  ChemComp-PIO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 24.0 µm / Nominal defocus min: 6.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)