+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Phi-29 scaffolding protein bound to intermediate-state MCP | |||||||||
 Map data Map data | Phi-29 scaffolding protein bound to intermediate-state MCP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | bacteriophage / prohead / scaffold / HK97 fold / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationviral scaffold / viral procapsid / T=3 icosahedral viral capsid / virion assembly / DNA binding Similarity search - Function | |||||||||
| Biological species |   Bacillus phage phi29 (virus) Bacillus phage phi29 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Woodson ME / Morais MC / Jardine PJ / Scott SD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2025 Journal: Sci Adv / Year: 2025Title: Phi29 assembly intermediates reveal how scaffold interactions with capsid protein drive capsid construction and maturation Authors: Woodson M / Prokhorov NS / Scott SD / Zhao W / Zhang W / Choi KH / Jardine PJ / Morais MC | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28822.map.gz emd_28822.map.gz | 26.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28822-v30.xml emd-28822-v30.xml emd-28822.xml emd-28822.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
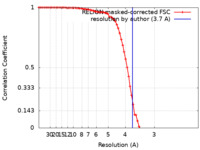
| FSC (resolution estimation) |  emd_28822_fsc.xml emd_28822_fsc.xml | 7.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_28822.png emd_28822.png | 182.2 KB | ||
| Filedesc metadata |  emd-28822.cif.gz emd-28822.cif.gz | 6.4 KB | ||
| Others |  emd_28822_half_map_1.map.gz emd_28822_half_map_1.map.gz emd_28822_half_map_2.map.gz emd_28822_half_map_2.map.gz | 29.4 MB 29.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28822 http://ftp.pdbj.org/pub/emdb/structures/EMD-28822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28822 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28822 | HTTPS FTP |
-Related structure data
| Related structure data |  8f2mMC  8f2nC  8f2oC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28822.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28822.map.gz / Format: CCP4 / Size: 38.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Phi-29 scaffolding protein bound to intermediate-state MCP | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.092 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half Map 1
| File | emd_28822_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
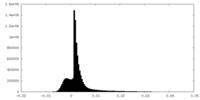
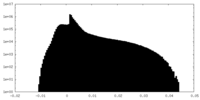
| Density Histograms |
-Half map: Half Map 2
| File | emd_28822_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
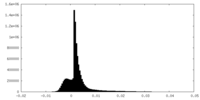
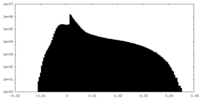
| Density Histograms |
- Sample components
Sample components
-Entire : Bacillus phage phi29
| Entire | Name:   Bacillus phage phi29 (virus) Bacillus phage phi29 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Bacillus phage phi29
| Supramolecule | Name: Bacillus phage phi29 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2884424 / Sci species name: Bacillus phage phi29 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 35.0 Å / T number (triangulation number): 3 |
-Macromolecule #1: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Bacillus phage phi29 (virus) Bacillus phage phi29 (virus) |
| Molecular weight | Theoretical: 49.894906 KDa |
| Recombinant expression | Organism:  Escherichia phage EcSzw-2 (virus) Escherichia phage EcSzw-2 (virus) |
| Sequence | String: MRITFNDVKT SLGITESYDI VNAIRNSQGD NFKSYVPLAT ANNVAEVGAG ILINQTVQND FITSLVDRIG LVVIRQVSLN NPLKKFKKG QIPLGRTIEE IYTDITKEKQ YDAEEAEQKV FEREMPNVKT LFHERNRQGF YHQTIQDDSL KTAFVSWGNF E SFVSSIIN ...String: MRITFNDVKT SLGITESYDI VNAIRNSQGD NFKSYVPLAT ANNVAEVGAG ILINQTVQND FITSLVDRIG LVVIRQVSLN NPLKKFKKG QIPLGRTIEE IYTDITKEKQ YDAEEAEQKV FEREMPNVKT LFHERNRQGF YHQTIQDDSL KTAFVSWGNF E SFVSSIIN AIYNSAEVDE YEYMKLLVDN YYSKGLFTTV KIDEPTSSTG ALTEFVKKMR ATARKLTLPQ GSRDWNSMAV RT RSYMEDL HLIIDADLEA ELDVDVLAKA FNMNRTDFLG NVTVIDGFAS TGLEAVLVDK DWFMVYDNLH KMETVRNPRG LYW NYYYHV WQTLSVSRFA NAVAFVSGDV PAVTQVIVSP NIAAVKQGGQ QQFTAYVRAT NAKDHKVVWS VEGGSTGTAI TGDG LLSVS GNEDNQLTVK ATVDIGTEDK PKLVVGEAVV SIRPNNASGG AQA UniProtKB: Major capsid protein |
-Macromolecule #2: Capsid assembly scaffolding protein
| Macromolecule | Name: Capsid assembly scaffolding protein / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Bacillus phage phi29 (virus) Bacillus phage phi29 (virus) |
| Molecular weight | Theoretical: 11.282529 KDa |
| Recombinant expression | Organism:  Escherichia phage EcSzw-2 (virus) Escherichia phage EcSzw-2 (virus) |
| Sequence | String: MPLKPEEHED ILNKLLDPEL AQSERTEALQ QLRVNYGSFV SEYNDLTKSH EKLAAEKDDL IVSNSKLFRQ IGLTDKQEED HKKADISET ITIEDLEAK UniProtKB: Capsid assembly scaffolding protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 1 / Number real images: 5593 / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8f2m: |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)