+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of substrate-free ClpX.ClpP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AAA+ protease / ClpXP / substrate-free / CHAPERONE | |||||||||
| Function / homology |  Function and homology information Function and homology informationHslUV protease complex / endopeptidase Clp / endopeptidase Clp complex / positive regulation of programmed cell death / response to temperature stimulus / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / proteasomal protein catabolic process / serine-type peptidase activity / : ...HslUV protease complex / endopeptidase Clp / endopeptidase Clp complex / positive regulation of programmed cell death / response to temperature stimulus / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / proteasomal protein catabolic process / serine-type peptidase activity / : / ATP-dependent protein folding chaperone / response to radiation / unfolded protein binding / peptidase activity / response to heat / ATPase binding / protein dimerization activity / serine-type endopeptidase activity / cell division / ATP hydrolysis activity / proteolysis / zinc ion binding / ATP binding / identical protein binding / membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.57 Å | |||||||||
 Authors Authors | Ghanbarpour A / Davis JH / Sauer RT | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure of substrate-free DNClpX.ClpP Authors: Ghanbarpour A / Cohen S / Davis JH / Sauer RT | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27952.map.gz emd_27952.map.gz | 32.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27952-v30.xml emd-27952-v30.xml emd-27952.xml emd-27952.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27952_fsc.xml emd_27952_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27952.png emd_27952.png | 52 KB | ||
| Masks |  emd_27952_msk_1.map emd_27952_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27952.cif.gz emd-27952.cif.gz | 5.8 KB | ||
| Others |  emd_27952_half_map_1.map.gz emd_27952_half_map_1.map.gz emd_27952_half_map_2.map.gz emd_27952_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27952 http://ftp.pdbj.org/pub/emdb/structures/EMD-27952 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27952 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27952 | HTTPS FTP |
-Related structure data
| Related structure data |  8e91MC  8e7vC  8e8qC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27952.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27952.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.124 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27952_msk_1.map emd_27952_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
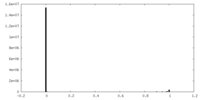
| Density Histograms |
-Half map: #1
| File | emd_27952_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
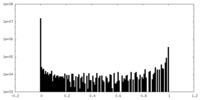
| Density Histograms |
-Half map: #2
| File | emd_27952_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
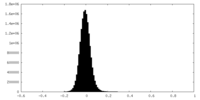
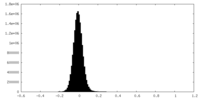
| Density Histograms |
- Sample components
Sample components
-Entire : ClpXP AAA protease
| Entire | Name: ClpXP AAA protease |
|---|---|
| Components |
|
-Supramolecule #1: ClpXP AAA protease
| Supramolecule | Name: ClpXP AAA protease / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ATP-dependent Clp protease ATP-binding subunit ClpX
| Macromolecule | Name: ATP-dependent Clp protease ATP-binding subunit ClpX / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 46.414848 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTDKRKDGSG KLLYCSFCGK SQHEVRKLIA GPSVYICDEC VDLCNDIIRE EIKEVAPHRE RSALPTPHEI RNHLDDYVIG QEQAKKVLA VAVYNHYKRL RNGDTSNGVE LGKSNILLIG PTGSGKTLLA ETLARLLDVP FTMADATTLT EAGYVGEDVE N IIQKLLQK ...String: MTDKRKDGSG KLLYCSFCGK SQHEVRKLIA GPSVYICDEC VDLCNDIIRE EIKEVAPHRE RSALPTPHEI RNHLDDYVIG QEQAKKVLA VAVYNHYKRL RNGDTSNGVE LGKSNILLIG PTGSGKTLLA ETLARLLDVP FTMADATTLT EAGYVGEDVE N IIQKLLQK CDYDVQKAQR GIVYIDEIDK ISRKSDNPSI TRDVSGEGVQ QALLKLIEGT VAAVPPQGGR KHPQQEFLQV DT SKILFIC GGAFAGLDKV ISHRVETGSG IGFGATVKAK SDKASEGELL AQVEPEDLIK FGLIPEFIGR LPVVATLNEL SEE ALIQIL KEPKNALTKQ YQALFNLEGV DLEFRDEALD AIAKKAMARK TGARGLRSIV EAALLDTMYD LPSMEDVEKV VIDE SVIDG QSEPLLIYGK PEAQQASGE UniProtKB: ATP-dependent Clp protease ATP-binding subunit ClpX |
-Macromolecule #2: ATP-dependent Clp protease proteolytic subunit
| Macromolecule | Name: ATP-dependent Clp protease proteolytic subunit / type: protein_or_peptide / ID: 2 / Number of copies: 14 / Enantiomer: LEVO / EC number: endopeptidase Clp |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.468869 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LVPMVIEQTS RGERSFDIYS RLLKERVIFL TGQVEDHMAN LIVAQMLFLE AENPEKDIYL YINSPGGVIT AGMSIYDTMQ FIKPDVSTI CMGQAASMGA FLLTAGAKGK RFCLPNSRVM IHQPLGGYQG QATDIEIHAR EILKVKGRMN ELMALHTGQS L EQIERDTE ...String: LVPMVIEQTS RGERSFDIYS RLLKERVIFL TGQVEDHMAN LIVAQMLFLE AENPEKDIYL YINSPGGVIT AGMSIYDTMQ FIKPDVSTI CMGQAASMGA FLLTAGAKGK RFCLPNSRVM IHQPLGGYQG QATDIEIHAR EILKVKGRMN ELMALHTGQS L EQIERDTE RDRFLSAPEA VEYGLVDSIL THRNENLYFQ SLEHHHHHH UniProtKB: ATP-dependent Clp protease proteolytic subunit |
-Macromolecule #3: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER / type: ligand / ID: 3 / Number of copies: 4 / Formula: AGS |
|---|---|
| Molecular weight | Theoretical: 523.247 Da |
| Chemical component information |  ChemComp-AGS: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 2 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 52.67 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.75 µm / Nominal defocus min: 0.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)