+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the PEAK3 pseudokinase homodimer | ||||||||||||
 Map data Map data | Reconstruction of the PEAK3 pseudokinase homodimer filtered to 4.9A and sharpened with -105 bFactor. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | complex / pseudokinase / kinase / adapter / SIGNALING PROTEIN | ||||||||||||
| Function / homology | : / regulation of cell motility / regulation of actin cytoskeleton organization / regulation of cell shape / actin cytoskeleton / protein kinase activity / focal adhesion / Protein PEAK3 Function and homology information Function and homology information | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
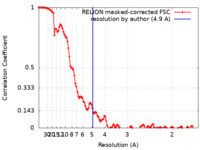
| Method | single particle reconstruction / cryo EM / Resolution: 4.9 Å | ||||||||||||
 Authors Authors | Torosyan H / Paul M / Jura N / Verba KA | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural insights into regulation of the PEAK3 pseudokinase scaffold by 14-3-3. Authors: Hayarpi Torosyan / Michael D Paul / Antoine Forget / Megan Lo / Devan Diwanji / Krzysztof Pawłowski / Nevan J Krogan / Natalia Jura / Kliment A Verba /   Abstract: PEAK pseudokinases are molecular scaffolds which dimerize to regulate cell migration, morphology, and proliferation, as well as cancer progression. The mechanistic role dimerization plays in PEAK ...PEAK pseudokinases are molecular scaffolds which dimerize to regulate cell migration, morphology, and proliferation, as well as cancer progression. The mechanistic role dimerization plays in PEAK scaffolding remains unclear, as there are no structures of PEAKs in complex with their interactors. Here, we report the cryo-EM structure of dimeric PEAK3 in complex with an endogenous 14-3-3 heterodimer. Our structure reveals an asymmetric binding mode between PEAK3 and 14-3-3 stabilized by one pseudokinase domain and the SHED domain of the PEAK3 dimer. The binding interface contains a canonical phosphosite-dependent primary interaction and a unique secondary interaction not observed in previous structures of 14-3-3/client complexes. Additionally, we show that PKD regulates PEAK3/14-3-3 binding, which when prevented leads to PEAK3 nuclear enrichment and distinct protein-protein interactions. Altogether, our data demonstrate that PEAK3 dimerization forms an unusual secondary interface for 14-3-3 binding, facilitating 14-3-3 regulation of PEAK3 localization and interactome diversity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27684.map.gz emd_27684.map.gz | 7.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27684-v30.xml emd-27684-v30.xml emd-27684.xml emd-27684.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27684_fsc.xml emd_27684_fsc.xml | 11.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_27684.png emd_27684.png | 60.1 KB | ||
| Masks |  emd_27684_msk_1.map emd_27684_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27684.cif.gz emd-27684.cif.gz | 6.2 KB | ||
| Others |  emd_27684_half_map_1.map.gz emd_27684_half_map_1.map.gz emd_27684_half_map_2.map.gz emd_27684_half_map_2.map.gz | 98.3 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27684 http://ftp.pdbj.org/pub/emdb/structures/EMD-27684 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27684 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27684 | HTTPS FTP |
-Related structure data
| Related structure data |  8ds6MC  8dp5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27684.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27684.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
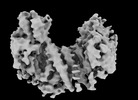
| Annotation | Reconstruction of the PEAK3 pseudokinase homodimer filtered to 4.9A and sharpened with -105 bFactor. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27684_msk_1.map emd_27684_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
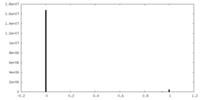
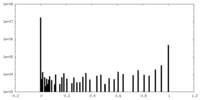
| Density Histograms |
-Half map: unfiltered, unsharpened half maps
| File | emd_27684_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered, unsharpened half maps | ||||||||||||
| Projections & Slices |
| ||||||||||||
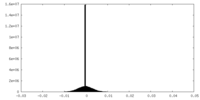
| Density Histograms |
-Half map: unfiltered, unsharpened half maps
| File | emd_27684_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered, unsharpened half maps | ||||||||||||
| Projections & Slices |
| ||||||||||||
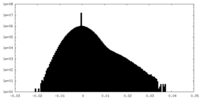
| Density Histograms |
- Sample components
Sample components
-Entire : PEAK3 pseudokinase homodimer
| Entire | Name: PEAK3 pseudokinase homodimer |
|---|---|
| Components |
|
-Supramolecule #1: PEAK3 pseudokinase homodimer
| Supramolecule | Name: PEAK3 pseudokinase homodimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 104.58 KDa |
-Macromolecule #1: Protein PEAK3
| Macromolecule | Name: Protein PEAK3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 52.357031 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSSPEPPTEP PEPDNPTWST QPTYSNLGQI RAHLLPSKAC RLRTPGSLST NPEPLPPPLP KKILTRTQSL PTRRTLHPSS IQVQPPRRP FLGSHSVDKS QAAVGPACLP AELTFGPADA PLGLSLRDLH SPEAVHTALA ARQLQGLRTI YARLRARLMG G HPGPCHPG ...String: MSSPEPPTEP PEPDNPTWST QPTYSNLGQI RAHLLPSKAC RLRTPGSLST NPEPLPPPLP KKILTRTQSL PTRRTLHPSS IQVQPPRRP FLGSHSVDKS QAAVGPACLP AELTFGPADA PLGLSLRDLH SPEAVHTALA ARQLQGLRTI YARLRARLMG G HPGPCHPG HSFRLLDSSP CAESGDALYY RVVRAHEDAW HILVAKVPKP GADVPHPWGL ELQASLSPHF NLQGLCGLVP EG TLPGAPW RGAVALAAEV PERTVAQWLA EACTQPPEEF VWAVALLLLQ LSAALKFLEA WGAALVELRP ENLLLVAPRG CAT TGPPRL LLTDFGRVCL QPPGPPGSPG PHAPQLGSLL RALLSLAAPS TTPLAAGLEL LAAQLTRLRP SASRTRGALQ ALLW GPGPE LRGRGAPLGP WLRALGPWLR VRRGLLVLRL AERAAGGEAP SLEDWLCCEY LAEATESSMG QALALLWDLE GGGGA DYKD DDDKGPV UniProtKB: Protein PEAK3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: A final concentration of 0.1% of Octyl-beta-Glucoside (C14H28O6) was added to the sample before freezing. | |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278.15 K / Instrument: FEI VITROBOT MARK IV / Details: blot time = 7s blot force = 4. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Average electron dose: 69.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)