[English] 日本語
 Yorodumi
Yorodumi- EMDB-27431: Cryo-EM structure of Saccharomyces cerevisiae Succinyl-CoA:acetat... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Saccharomyces cerevisiae Succinyl-CoA:acetate CoA-transferase (Ach1p) | |||||||||
 Map data Map data | map sharp | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ach1p / Succinyl-CoA / acetate CoA-transferase / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationacetyl-CoA hydrolase / acetyl-CoA hydrolase activity / acetate CoA-transferase activity / acetate metabolic process / mitochondrion / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
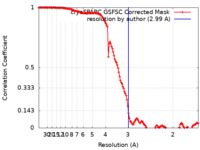
| Method | single particle reconstruction / cryo EM / Resolution: 2.99 Å | |||||||||
 Authors Authors | Godoy AS / Song Y / Cheruvara H / Quigley A / Oliva G | |||||||||
| Funding support |  Brazil, 1 items Brazil, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of Saccharomyces cerevisiae cytochrome c oxidase (Complex IV) extracted in lipid nanodiscs Authors: Godoy AS / Song Y / Cheruvara H / Quigley A / Oliva G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27431.map.gz emd_27431.map.gz | 229.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27431-v30.xml emd-27431-v30.xml emd-27431.xml emd-27431.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27431_fsc.xml emd_27431_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_27431.png emd_27431.png | 83.9 KB | ||
| Masks |  emd_27431_msk_1.map emd_27431_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27431.cif.gz emd-27431.cif.gz | 5.7 KB | ||
| Others |  emd_27431_additional_1.map.gz emd_27431_additional_1.map.gz emd_27431_half_map_1.map.gz emd_27431_half_map_1.map.gz emd_27431_half_map_2.map.gz emd_27431_half_map_2.map.gz | 122 MB 226.2 MB 226.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27431 http://ftp.pdbj.org/pub/emdb/structures/EMD-27431 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27431 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27431 | HTTPS FTP |
-Validation report
| Summary document |  emd_27431_validation.pdf.gz emd_27431_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27431_full_validation.pdf.gz emd_27431_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_27431_validation.xml.gz emd_27431_validation.xml.gz | 21.8 KB | Display | |
| Data in CIF |  emd_27431_validation.cif.gz emd_27431_validation.cif.gz | 27.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27431 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27431 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27431 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27431 | HTTPS FTP |
-Related structure data
| Related structure data |  8dh7MC  8dh6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27431.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27431.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map sharp | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.831 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27431_msk_1.map emd_27431_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: map
| File | emd_27431_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfA
| File | emd_27431_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfA | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27431_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Succinyl-CoA:acetate CoA-transferase
| Entire | Name: Succinyl-CoA:acetate CoA-transferase |
|---|---|
| Components |
|
-Supramolecule #1: Succinyl-CoA:acetate CoA-transferase
| Supramolecule | Name: Succinyl-CoA:acetate CoA-transferase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Acetyl-CoA hydrolase
| Macromolecule | Name: Acetyl-CoA hydrolase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: acetyl-CoA hydrolase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 58.791785 KDa |
| Sequence | String: MTISNLLKQR VRYAPYLKKV KEAHELIPLF KNGQYLGWSG FTGVGTPKAV PEALIDHVEK NNLQGKLRFN LFVGASAGPE ENRWAEHDM IIKRAPHQVG KPIAKAINQG RIEFFDKHLS MFPQDLTYGF YTRERKDNKI LDYTIIEATA IKEDGSIVPG P SVGGSPEF ...String: MTISNLLKQR VRYAPYLKKV KEAHELIPLF KNGQYLGWSG FTGVGTPKAV PEALIDHVEK NNLQGKLRFN LFVGASAGPE ENRWAEHDM IIKRAPHQVG KPIAKAINQG RIEFFDKHLS MFPQDLTYGF YTRERKDNKI LDYTIIEATA IKEDGSIVPG P SVGGSPEF ITVSDKVIIE VNTATPSFEG IHDIDMPVNP PFRKPYPYLK VDDKCGVDSI PVDPEKVVAI VESTMRDQVP PN TPSDDMS RAIAGHLVEF FRNEVKHGRL PENLLPLQSG IGNIANAVIE GLAGAQFKHL TVWTEVLQDS FLDLFENGSL DYA TATSVR LTEKGFDRAF ANWENFKHRL CLRSQVVSNN PEMIRRLGVI AMNTPVEVDI YAHANSTNVN GSRMLNGLGG SADF LRNAK LSIMHAPSAR PTKVDPTGIS TIVPMASHVD QTEHDLDILV TDQGLADLRG LSPKERAREI INKCAHPDYQ ALLTD YLDR AEHYAKKHNC LHEPHMLKNA FKFHTNLAEK GTMKVDSWEP VD UniProtKB: Acetyl-CoA hydrolase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 0.989314222 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.1 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)