[English] 日本語
 Yorodumi
Yorodumi- EMDB-27256: Human DNA polymerase alpha/primase elongation complex I bound to ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human DNA polymerase alpha/primase elongation complex I bound to primer/template | |||||||||
 Map data Map data | Unsharpened and flipped and resampled map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA replication / human DNA polymerase alpha/primase / human primosome / elongation complex / REPLICATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of DNA primase activity / DNA replication initiation / DNA/RNA hybrid binding / Telomere C-strand synthesis initiation / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / regulation of type I interferon production / Polymerase switching / alpha DNA polymerase:primase complex / Processive synthesis on the lagging strand / Removal of the Flap Intermediate ...positive regulation of DNA primase activity / DNA replication initiation / DNA/RNA hybrid binding / Telomere C-strand synthesis initiation / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / regulation of type I interferon production / Polymerase switching / alpha DNA polymerase:primase complex / Processive synthesis on the lagging strand / Removal of the Flap Intermediate / Polymerase switching on the C-strand of the telomere / primosome complex / DNA replication, synthesis of primer / lagging strand elongation / mitotic DNA replication initiation / DNA strand elongation involved in DNA replication / DNA synthesis involved in DNA repair / G1/S-Specific Transcription / leading strand elongation / DNA replication origin binding / Activation of the pre-replicative complex / DNA replication initiation / Defective pyroptosis / double-strand break repair via nonhomologous end joining / nuclear matrix / nuclear envelope / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / DNA replication / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / DNA repair / nucleotide binding / chromatin binding / chromatin / nucleolus / protein kinase binding / DNA binding / zinc ion binding / nucleoplasm / nucleus / metal ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
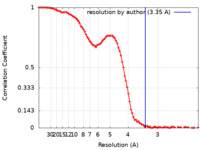
| Method | single particle reconstruction / cryo EM / Resolution: 3.35 Å | |||||||||
 Authors Authors | He Q / Baranovskiy A / Lim C / Tahirov T | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structures of human primosome elongation complexes. Authors: Qixiang He / Andrey G Baranovskiy / Lucia M Morstadt / Alisa E Lisova / Nigar D Babayeva / Benjamin L Lusk / Ci Ji Lim / Tahir H Tahirov /  Abstract: The synthesis of RNA-DNA primer by primosome requires coordination between primase and DNA polymerase α subunits, which is accompanied by unknown architectural rearrangements of multiple domains. ...The synthesis of RNA-DNA primer by primosome requires coordination between primase and DNA polymerase α subunits, which is accompanied by unknown architectural rearrangements of multiple domains. Using cryogenic electron microscopy, we solved a 3.6 Å human primosome structure caught at an early stage of RNA primer elongation with deoxynucleotides. The structure confirms a long-standing role of primase large subunit and reveals new insights into how primosome is limited to synthesizing short RNA-DNA primers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27256.map.gz emd_27256.map.gz | 30.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27256-v30.xml emd-27256-v30.xml emd-27256.xml emd-27256.xml | 23.6 KB 23.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27256_fsc.xml emd_27256_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27256.png emd_27256.png | 96 KB | ||
| Filedesc metadata |  emd-27256.cif.gz emd-27256.cif.gz | 7.8 KB | ||
| Others |  emd_27256_half_map_1.map.gz emd_27256_half_map_1.map.gz emd_27256_half_map_2.map.gz emd_27256_half_map_2.map.gz | 48.8 MB 48.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27256 http://ftp.pdbj.org/pub/emdb/structures/EMD-27256 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27256 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27256 | HTTPS FTP |
-Validation report
| Summary document |  emd_27256_validation.pdf.gz emd_27256_validation.pdf.gz | 723.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27256_full_validation.pdf.gz emd_27256_full_validation.pdf.gz | 722.7 KB | Display | |
| Data in XML |  emd_27256_validation.xml.gz emd_27256_validation.xml.gz | 15.7 KB | Display | |
| Data in CIF |  emd_27256_validation.cif.gz emd_27256_validation.cif.gz | 21 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27256 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27256 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27256 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27256 | HTTPS FTP |
-Related structure data
| Related structure data |  8d96MC  8d9dC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27256.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27256.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened and flipped and resampled map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map B
| File | emd_27256_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_27256_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Elongation complex I of human DNA polymerase alpha/primase bound ...
| Entire | Name: Elongation complex I of human DNA polymerase alpha/primase bound to RNA-DNA primer and DNA template |
|---|---|
| Components |
|
-Supramolecule #1: Elongation complex I of human DNA polymerase alpha/primase bound ...
| Supramolecule | Name: Elongation complex I of human DNA polymerase alpha/primase bound to RNA-DNA primer and DNA template type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: Smaller elongation complex solved using cryo-EM single-particle analysis |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 340 KDa |
-Macromolecule #1: DNA primase large subunit
| Macromolecule | Name: DNA primase large subunit / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.890918 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEFSGRKWRK LRLAGDQRNA SYPHCLQFYL QPPSENISLI EFENLAIDRV KLLKSVENLG VSYVKGTEQY QSKLESELRK LKFSYRENL EDEYEPRRRD HISHFILRLA YCQSEELRRW FIQQEMDLLR FRFSILPKDK IQDFLKDSQL QFEAISDEEK T LREQEIVA ...String: MEFSGRKWRK LRLAGDQRNA SYPHCLQFYL QPPSENISLI EFENLAIDRV KLLKSVENLG VSYVKGTEQY QSKLESELRK LKFSYRENL EDEYEPRRRD HISHFILRLA YCQSEELRRW FIQQEMDLLR FRFSILPKDK IQDFLKDSQL QFEAISDEEK T LREQEIVA SSPSLSGLKL GFESIYKIPF ADALDLFRGR KVYLEDGFAY VPLKDIVAII LNEFRAKLSK ALALTARSLP AV QSDERLQ PLLNHLSHSY TGQDYSTQGN VGKISLDQID LLSTKSFPPC MRQLHKALRE NHHLRHGGRM QYGLFLKGIG LTL EQALQF WKQEFIKGKM DPDKFDKGYS YNIRHSFGKE GKRTDYTPFS CLKIILSNPP SQGDYHGCPF RHSDPELLKQ KLQS YKISP GGISQILDLV KGTHYQVACQ KYFEMIHNVD DCGFSLNHPN QFFCESQRIL NGGKDIKKEP IQPETPQPKP SVQKT KDAS SALASLNSSL EMDMEGLEDY FSEDS UniProtKB: DNA primase large subunit |
-Macromolecule #2: DNA polymerase alpha catalytic subunit
| Macromolecule | Name: DNA polymerase alpha catalytic subunit / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 166.131094 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAPVHGDDSL SDSGSFVSSR ARREKKSKKG RQEALERLKK AKAGEKYKYE VEDFTGVYEE VDEEQYSKLV QARQDDDWIV DDDGIGYVE DGREIFDDDL EDDALDADEK GKDGKARNKD KRNVKKLAVT KPNNIKSMFI ACAGKKTADK AVDLSKDGLL G DILQDLNT ...String: MAPVHGDDSL SDSGSFVSSR ARREKKSKKG RQEALERLKK AKAGEKYKYE VEDFTGVYEE VDEEQYSKLV QARQDDDWIV DDDGIGYVE DGREIFDDDL EDDALDADEK GKDGKARNKD KRNVKKLAVT KPNNIKSMFI ACAGKKTADK AVDLSKDGLL G DILQDLNT ETPQITPPPV MILKKKRSIG ASPNPFSVHT ATAVPSGKIA SPVSRKEPPL TPVPLKRAEF AGDDVQVEST EE EQESGAM EFEDGDFDEP MEVEEVDLEP MAAKAWDKES EPAEEVKQEA DSGKGTVSYL GSFLPDVSCW DIDQEGDSSF SVQ EVQVDS SHLPLVKGAD EEQVFHFYWL DAYEDQYNQP GVVFLFGKVW IESAETHVSC CVMVKNIERT LYFLPREMKI DLNT GKETG TPISMKDVYE EFDEKIATKY KIMKFKSKPV EKNYAFEIPD VPEKSEYLEV KYSAEMPQLP QDLKGETFSH VFGTN TSSL ELFLMNRKIK GPCWLEVKSP QLLNQPVSWC KVEAMALKPD LVNVIKDVSP PPLVVMAFSM KTMQNAKNHQ NEIIAM AAL VHHSFALDKA APKPPFQSHF CVVSKPKDCI FPYAFKEVIE KKNVKVEVAA TERTLLGFFL AKVHKIDPDI IVGHNIY GF ELEVLLQRIN VCKAPHWSKI GRLKRSNMPK LGGRSGFGER NATCGRMICD VEISAKELIR CKSYHLSELV QQILKTER V VIPMENIQNM YSESSQLLYL LEHTWKDAKF ILQIMCELNV LPLALQITNI AGNIMSRTLM GGRSERNEFL LLHAFYENN YIVPDKQIFR KPQQKLGDED EEIDGDTNKY KKGRKKAAYA GGLVLDPKVG FYDKFILLLD FNSLYPSIIQ EFNICFTTVQ RVASEAQKV TEDGEQEQIP ELPDPSLEMG ILPREIRKLV ERRKQVKQLM KQQDLNPDLI LQYDIRQKAL KLTANSMYGC L GFSYSRFY AKPLAALVTY KGREILMHTK EMVQKMNLEV IYGDTDSIMI NTNSTNLEEV FKLGNKVKSE VNKLYKLLEI DI DGVFKSL LLLKKKKYAA LVVEPTSDGN YVTKQELKGL DIVRRDWCDL AKDTGNFVIG QILSDQSRDT IVENIQKRLI EIG ENVLNG SVPVSQFEIN KALTKDPQDY PDKKSLPHVH VALWINSQGG RKVKAGDTVS YVICQDGSNL TASQRAYAPE QLQK QDNLT IDTQYYLAQQ IHPVVARICE PIDGIDAVLI ATWLGLDPTQ FRVHHYHKDE ENDALLGGPA QLTDEEKYRD CERFK CPCP TCGTENIYDN VFDGSGTDME PSLYRCSNID CKASPLTFTV QLSNKLIMDI RRFIKKYYDG WLICEEPTCR NRTRHL PLQ FSRTGPLCPA CMKATLQPEY SDKSLYTQLC FYRYIFDAEC ALEKLTTDHE KDKLKKQFFT PKVLQDYRKL KNTAEQF LS RSGYSEVNLS KLFAGCAVKS UniProtKB: DNA polymerase alpha catalytic subunit |
-Macromolecule #3: DNA/RNA (5'-GTP)-R(P*GP*CP*GP*GP*CP*AP*CP*G)-D(P*AP*CP*C)-3')
| Macromolecule | Name: DNA/RNA (5'-GTP)-R(P*GP*CP*GP*GP*CP*AP*CP*G)-D(P*AP*CP*C)-3') type: other / ID: 3 / Number of copies: 1 Classification: polydeoxyribonucleotide/polyribonucleotide hybrid |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.977352 KDa |
| Sequence | String: (GTP)GCGGCACG(DA) (DC)(DC) |
-Macromolecule #4: DNA (5'-D(*AP*TP*AP*AP*TP*GP*GP*TP*CP*GP*TP*GP*CP*CP*GP*CP*CP*AP*...
| Macromolecule | Name: DNA (5'-D(*AP*TP*AP*AP*TP*GP*GP*TP*CP*GP*TP*GP*CP*CP*GP*CP*CP*AP*AP*TP*AP*A)-3') type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.760392 KDa |
| Sequence | String: (DA)(DT)(DA)(DA)(DT)(DG)(DG)(DT)(DC)(DG) (DT)(DG)(DC)(DC)(DG)(DC)(DC)(DA)(DA)(DT) (DA)(DA) |
-Macromolecule #5: IRON/SULFUR CLUSTER
| Macromolecule | Name: IRON/SULFUR CLUSTER / type: ligand / ID: 5 / Number of copies: 1 / Formula: SF4 |
|---|---|
| Molecular weight | Theoretical: 351.64 Da |
| Chemical component information |  ChemComp-FS1: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #7: 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE
| Macromolecule | Name: 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE / type: ligand / ID: 7 / Number of copies: 1 / Formula: DTP |
|---|---|
| Molecular weight | Theoretical: 491.182 Da |
| Chemical component information |  ChemComp-DTP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: CHAPSO is made fresh at 80 mM before added to the sample at a final concentration of 4-8 mM immediately before vitrification. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 13243 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)