[English] 日本語
 Yorodumi
Yorodumi- EMDB-27142: Structure of Acidothermus cellulolyticus Cas9 ternary complex (Cl... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
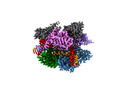
| Title | Structure of Acidothermus cellulolyticus Cas9 ternary complex (Cleavage Intermediate 1) | |||||||||
 Map data Map data | AcCas9 Cleavage intermediate 1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cas9 / AcCas9 / Crispr / RNA BINDING PROTEIN / RNA BINDING PROTEIN-DNA-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationendonuclease activity / defense response to virus / hydrolase activity / DNA binding / RNA binding / zinc ion binding Similarity search - Function | |||||||||
| Biological species |  Acidothermus cellulolyticus 11B (bacteria) / synthetic construct (others) Acidothermus cellulolyticus 11B (bacteria) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.21 Å | |||||||||
 Authors Authors | Rai J / Das A / Li H | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Catal / Year: 2023 Journal: Nat Catal / Year: 2023Title: Coupled catalytic states and the role of metal coordination in Cas9. Authors: Anuska Das / Jay Rai / Mitchell O Roth / Yuerong Shu / Megan L Medina / Mackenzie R Barakat / Hong Li /  Abstract: Controlling the activity of the CRISPR-Cas9 system is essential to its safe adoption for clinical and research applications. Although the conformational dynamics of Cas9 are known to control its ...Controlling the activity of the CRISPR-Cas9 system is essential to its safe adoption for clinical and research applications. Although the conformational dynamics of Cas9 are known to control its enzymatic activity, details of how Cas9 influences the catalytic processes at both nuclease domains remain elusive. Here we report five cryo-electron microscopy structures of the active Cas9 complex along the reaction path at 2.2-2.9 Å resolution. We observed that a large movement in one nuclease domain, triggered by the cognate DNA, results in noticeable changes in the active site of the other domain that is required for metal coordination and catalysis. Furthermore, the conformations synchronize the reaction intermediates, enabling coupled cutting of the two DNA strands. Consistent with the roles of conformations in organizing the active sites, adjustments to the metal-coordination residues lead to altered metal specificity of Cas9 and commonly used Cas9 in cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27142.map.gz emd_27142.map.gz | 100.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27142-v30.xml emd-27142-v30.xml emd-27142.xml emd-27142.xml | 24.7 KB 24.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27142.png emd_27142.png | 50.6 KB | ||
| Filedesc metadata |  emd-27142.cif.gz emd-27142.cif.gz | 7.6 KB | ||
| Others |  emd_27142_half_map_1.map.gz emd_27142_half_map_1.map.gz emd_27142_half_map_2.map.gz emd_27142_half_map_2.map.gz | 84.2 MB 84.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27142 http://ftp.pdbj.org/pub/emdb/structures/EMD-27142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27142 | HTTPS FTP |
-Related structure data
| Related structure data |  8d2lMC  8d2kC  8d2nC  8d2oC  8d2pC  8d2qC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27142.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27142.map.gz / Format: CCP4 / Size: 107.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AcCas9 Cleavage intermediate 1 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Halfmap1
| File | emd_27142_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap2
| File | emd_27142_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : CryoEM Structure of AceCas9 (Cleavage intermediate 1)
+Supramolecule #1: CryoEM Structure of AceCas9 (Cleavage intermediate 1)
+Supramolecule #2: CRISPR-associated endonuclease, Csn1 family/RNA
+Supramolecule #3: DNA
+Macromolecule #1: CRISPR-associated endonuclease, Csn1 family
+Macromolecule #2: Single guide RNA (106-MER)
+Macromolecule #3: DNA target strand (5'-D(*AP*GP*CP*TP*TP*GP*GP*TP*GP*TP*AP*TP*A)-3')
+Macromolecule #4: DNA target strand (5'-D(P*CP*CP*AP*GP*GP*AP*TP*CP*TP*TP*GP*CP*CP*...
+Macromolecule #5: DNA non-target strand (5'-D(P*TP*AP*TP*AP*CP*AP*CP*CP*AP*AP*GP*CP...
+Macromolecule #6: DNA non-target strand (5'-D(P*AP*GP*A)-3')
+Macromolecule #7: MAGNESIUM ION
+Macromolecule #8: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)